| [Molecules: 3] [Related articles/posters: 067 115 032 109 114 ] |
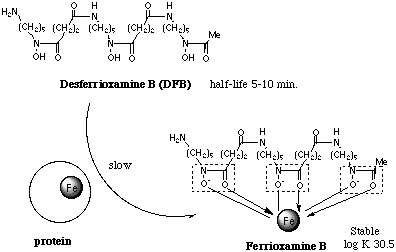
The kinetic studies of iron removal from transferrin have been reported, and an interaction between chelators and transferrin on iron exchange process was suggested. The process was found to be catalysed by pyrophosphate, which induced the conformation change of the protein. Neiland and coworkers demonstrated that aerobactin can remove iron from transferrin in vitro, and they showed spectrophotometric evidence for ternary complex formation in the process of iron removal from the protein. They suggested that the rate of iron exchange between transferrin and aerobactin is controlled by the rate of the conformational change of transferrin and the rate of dissociation of the complex. Cowart and coworkers also proposed the formation of a complex between the protein and the chelator; the complexation promotes a conformation change of the protein to expose the metal site. On the other hand, Raymond and coworkers suggested that the rate of iron removal from transferrin strongly depends on anionic property of the chelators. Thus, it is likely that the kinetic behaviour of the chelators is closely related to not only its anionic property, but also the structural suitability to the protein. However, no paper concerning the relationship between the ligand structure including the chirality and the kinetic behaviour has been reported.
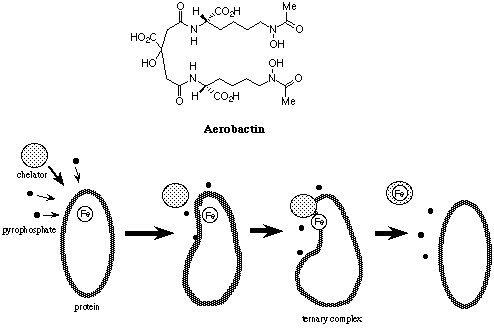
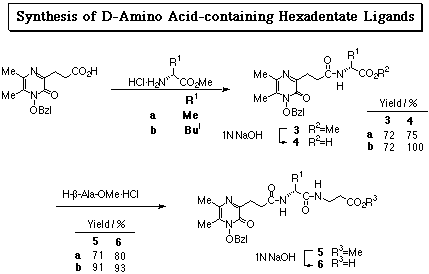
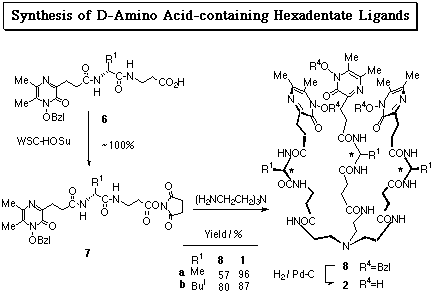
N-Hydroxyamide-containing pyrazines showed feasibility as iron sequestering agents under physiological conditions by virtue of their high water solubility and high acidity. It was also elucidated that chiral hexadenate chelators (1) composed of 1-hydroxy-5,6-dimethyl-2(1H)-pyrazinones, L-amino acid residues, and tris(2-aminoethyl)amine effectively removed iron from transferrin (J. Ohkanda and A. Katoh, J. Org. Chem., 1995, 60, 1583). As an attempt to explore the relationship between ligand structure and the iron removal efficiency, their enantiomers 2a, b were prepared.
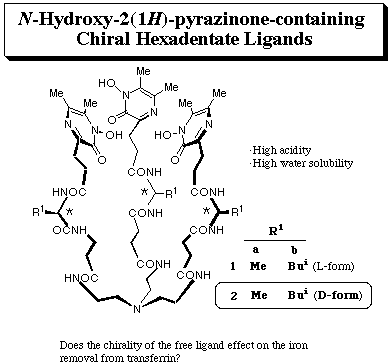
Iron(III)-chelating properties:
UV-VIS spectra of a 1:1 molar mixture of 2 and ferric ion in aqueous solutions showed characteristic LMCT band at ca. 450 nm (epsilon 3580 at 450 nm for 2a; epsilon 3010 at 455 nm for 2b), indicating the formation of intramolecular 1:1 iron complexes.
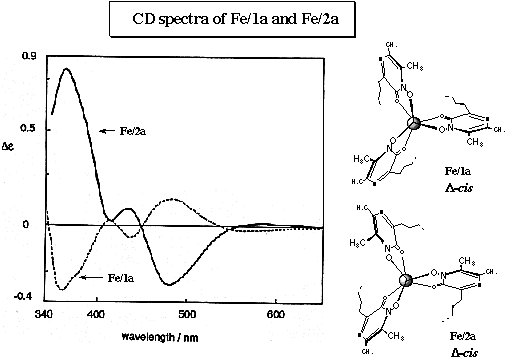
CD spectrum of the Fe/2a complex in an aqueous solution is shown in the next figure with the data of Fe/1a complex. The spectrum of Fe/2a complex has a negative band at 480 nm which is opposite to the band at 478 nm in Fe/1a complex. These bands arise from LMCT of the complexes and are therefore sensitive to the chirality at the metal centre. On the basis of these results, the absolute configuration in Fe/2a complex could be assigned as the d form (opposite to Fe/1a complex).

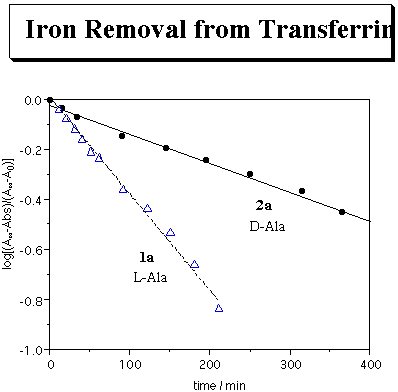
The iron removing ability of 2a from human diferric transferrin (TfFe 2.0) was evaluated at pH 7.4 by the same procedure as that for 1a. The plots of log[(A*-Abs)/(A*-A0)] as a function of time gave a linear relationship as shown in the next figure. It indicates that iron removal from transferrin by 2a proceeds with pseudo-first-order kinetics. The kobs was calculated from the slope of the straight line. The kinetic results are summarized in the Table together with the data of 1a, b. The rates of iron removal by 2a were greater than that by desferrioxamine B, even at a lower ligand concentration. It is noteworthy that the rates of iron removal by alanine residue-containing ligands were significantly affected by the chirality of the free ligands. The rate of iron removal by 2a was suppressed to only one quarter of 1a. On the other hand, no apparent difference was observed in the case of leucine residue-containing 2b and 1b.
| Ligand | [L]/[TfFe 2.0]a | kobs/10-3 min-1 | Fe removed (%) |
| 1a | 5 | 3.80 | 23 |
| 2a | 5 | 1.05 | 7 |
| 1b | 6 | 0.90 | 9 |
| 2b | 6 | 1.29 | 8 |
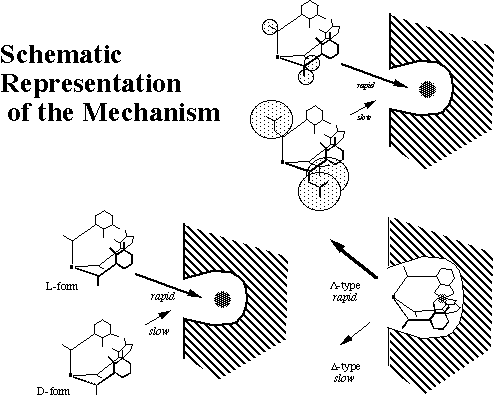
This is the first demonstration of the influence of the ligand's chirality on the rate of iron removal from transferrin. Although the detail remains obscure, a plausible mechanism is proposed as follows.
Ligand structural features influence the interaction between the protein and the ligand. A bulky substituent such as an Bui group may prevent access of the ligand to the exposed metal site of the protein. The iron removal process could be affected by the chirality of the ligand itself, or the absolute configuration (i.e. L and D) of the resulting iron complex of the ligand.