Origins of G-Quadruplex DNA
Chromosome
Our bodies are composed of more than a billion cells. Cells are continually dying and new cells are continually being formed. An identical copy of our hereditary material is found in the nucleus of each and every somatic cell. By viewing a nucleus under a microscope our chromosomes can actually be seen. There are 46 chromosomes present in the nucleus of each human cell, which are subsequently divided into 23 pairs of x-shaped bundles.
Fig 1. Chromosome when viewed under a microscope.
What are Chromosomes Made Of?
The arms of a chromosome are made up of long strands of DNA wound round in tight coils to form the chromosome. DNA is short for deoxyribonucleic acid, it is a special chemical that carries the hereditary information in almost all living things. The DNA tells the cell how to work and which particular characteristics it should have. A section of DNA that tells the cell to display a certain characteristic, or produce a certain particular chemical is called a gene. Each cell in your body contains 40,000 different genes. In a pair, each "arm" of the chromosome contains identical information along its length e.g. both "arms" contain the gene for eye colour, hair colour etc. The genes are arranged in the same order in both "arms". Each time a cell divides; each chromosome must be carefully replicated (copied).
The structure of DNA
Fig 2. Structure of a nucleotide
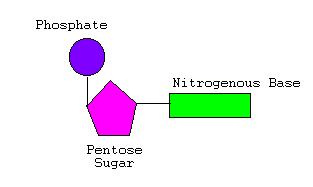
There are 4 different types of nitrogenous bases that can be found in the nucleotides of a DNA molecule. Each base is usually known by the first letter of its name;
A single strand of DNA is a long chain of nucleotides, each with one of these four different bases. This strand is represented by a sequence of these individual letters e.g. AGTCTTCAGGT. Usually two strands of nucleotides wrap around each other, giving DNA the appearance of a twisted ladder, called a double helix. The backbone chain of pentose sugar-phosphate links forms the struts of the ladder and is always on the outside. The rungs of the ladder are made up of base pairs, which are almost always found connected to each other. In a complete helix the Adenine(A) always lines up with Thymine(T) and the Guanine(G) with Cytosine(C). In these combinations the different bases fit together perfectly like a lock and key. This is termed the Watson-Crick base- pairing scheme.
Fig 3. Base Pairing in DNA Double Helix.
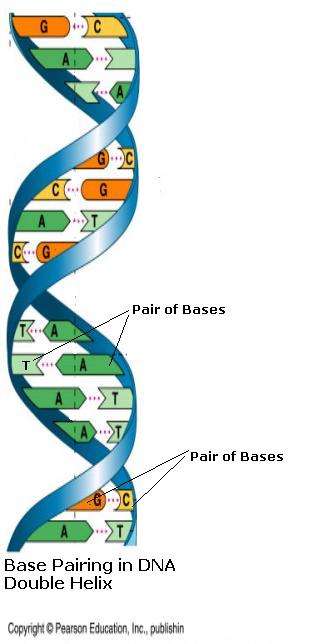
Chromosome Replication
DNA replication is when chromosomes duplicate themselves. The first step is to unwind their double helices into separate strands. As the double helix of DNA unwinds into two parent strands, the ends of the different bases are exposed. Due to the obligatory pairing of A-T and G-C, each parent strand becomes a template for copying a whole new DNA helix. Since the DNA structure can be rebuilt on both parent strands, two identical DNA helices are produced, each containing one original parent strand and one newly synthesized strand, called a complementary strand. Due to the nature of the mechanism via which DNA is replicated, one strand of the DNA is left with an incompletely replicated end. Without specialized means of maintaining chromosomes, this causes chromosome ends to shrink with each successive cell division.
Fig 4. Replication of DNA.
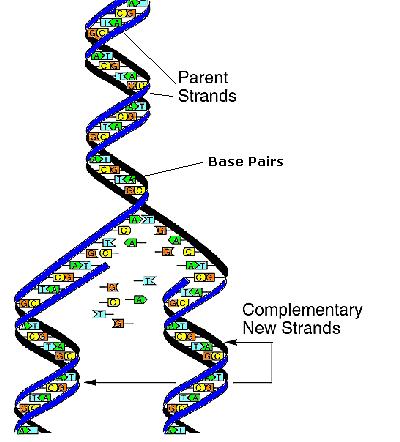
What is Cancer?
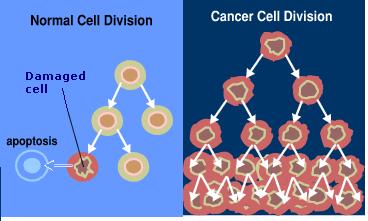
A cell needs to divide in order to preserve the organism that it supports. In normal cells, the rate of new cell growth is kept in balance with the rate at which old cells die. In cancer cells this balance is disrupted either by the loss of normal growth control or the loss of a cell's ability to undergo programmed cell death, known as apoptosis. Cancer cells originate when a cell does not replicate itself perfectly, this increases its chance of genetic mutation. These cancer cells and their offspring then multiply uncontrollably and can invade and damage nearby tissue.
Fig 5. Cell Division in Normal and Cancer Cells.
Protecting the ends of our Chromosomes
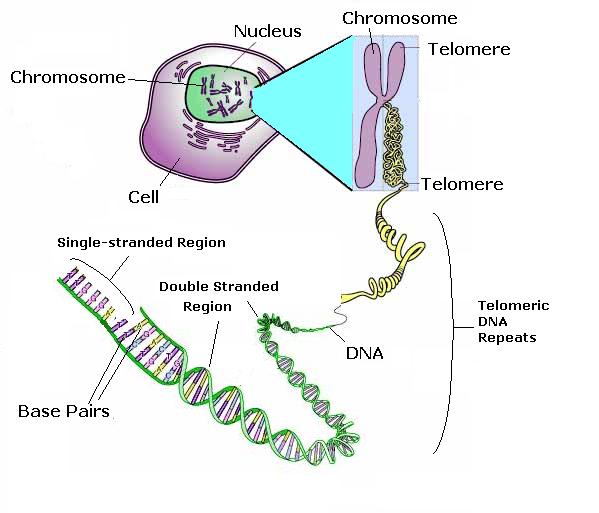
To prevent the loss or mutation of essential genes, the end of each chromosome is tied up by a special strand of DNA called a telomere. Telomeres cap our chromosomes preventing them from a wide range of otherwise catastrophic events and enable the DNA molecule to replicate. Telomeres therefore play an important role in maintaining the integrity of the DNA in our cells.
Fig 6. The telomere
Why do we need Telomeres?
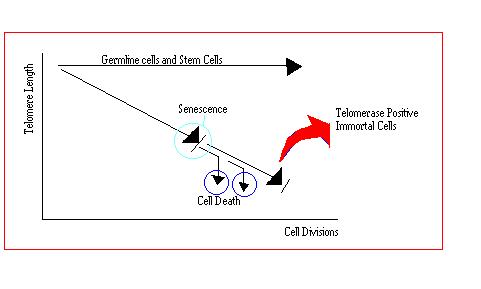
It is thought that the telomeres at the end of chromosomes may have evolved to prevent the unlimited growth of cells by limiting their life span. Evidence that average telomere length correlates with the lifespan of cell in culture, suggests that the growth of human cells is limited by their resource of telomeric DNA. In normal cells, mechanisms for maintaining telomere length are absent. So with each successive round of cell division, telomeres progressively decrease in length, loosing up to 200 base pairs of terminal DNA from the tips of their chromosomes with each division.[10] Since each chromosome possesses a telomere of finite length, the number of divisions a cell can undergo is limited by this store of telomeric DNA.[15] The more times a cell divides, the shorter the telomere becomes and ultimately after a limited number of cell divisions, the length of the telomere becomes so short that the cell ceases to divide. This non-dividing state is known as senescence. Almost all of these cells die due to the erosion of essential genes on at least one chromosome. The telomeres of certain cells do not shorten with progressive divisions; these cells are stem cells, germline cells and cancer cells.
Fig 7. Behaviour of Cells and their telomere lengths
Telomeres in Cancer Cells
Almost all tumour cells have markedly shortened telomeres in comparison to their counterparts in normal cells. This indicates that the initial formative stages of cancer development must involve the erosion of telomeres.[14] Another important characteristic to note is that telomeres in tumour cells are maintained at a constant length.
Catastrophic Events
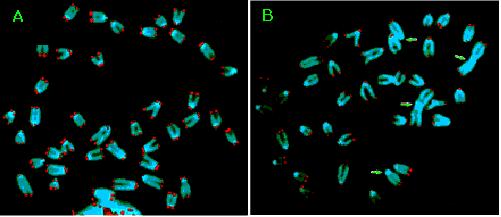
Fig 8. Fusion of Chromosomes
Replication of telomeres
The bulk of telomeric DNA has to be copied like the rest of the DNA in the chromosome. The replication of telomeres poses a special dilemma called the end-replication problem. This occurs because it is not possible to completely replicate the DNA through to the extreme end of the chromosome. Since the end region of the chromosome is where the telomere is located, it is the telomeric DNA that is lost during replication. If telomeres were not present, important genetic information would be lost with each cell division. As it is telomeres do not carry genetic information thus they act as a disposable part of DNA.
Structure of a Telomere
The DNA of human telomeres comprises of an extremely short and simple sequence of nucleotides, TTAGGG,[28,29] repeated over and over. An important characteristic to note is that the telomere strand is particularly rich in Guanine(G) bases.
Fig 9. Telomere G-rich Sequence
While most of the telomeric repeats exist in the double-stranded region of DNA, the extreme end of the telomere is single stranded. A crucial function of this region is the ability of the G-rich repeated DNA sequences to act as attachment points called binding sites. These binding sites are specifically designed for the attachment of particular proteins involved in the protection of the telomere end. The proteins work by both capping the telomere ends and assisting in the formation of more complex telomere structures. Ultimately the proteins contribute to the protection of the chromosome ends from the Catastrophic Events previously mentioned. The precise uses of some of the main proteins encountered are explained below.
Structure of a Telomere
The first protein to mention is Pot1, this stands for Protection Of Telomeres 1. The Pot1 protein has been found to bind to the single-stranded DNA and to actually cap the ends of the telomeres.
Two binding proteins have also been identified that bind to the double-stranded region of the telomere. The first of these proteins is TRF1, this stands for Telomeric Repeat Binding Factor 1. The TRF1 protein binds to the double-strand by recognizing the specific telomeric DNA sequence.
The second of these proteins is called TRF2 (Telomeric Repeat Binding Factor 2). TRF2 binds at the junction between the double and single stranded areas of DNA. This allows the extreme single-stranded end of the telomere to be folded-back and completely buried within the double stranded region, forming a complex structure called a t-loop.[13]
Fig 10 Formation of a t-loop
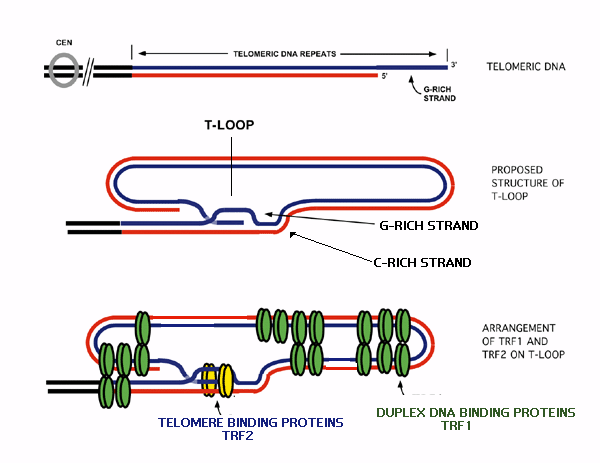
T-loops have been visualized in vitro. These t-loops have been proposed to exist in order to control telomere length. As the telomere lengthens a t-loop forms, blocking the action of the enzyme that extends the telomere. This prevents any further lengthening of the telomere. (The mechanism of this process is discussed later in Maintenance of Telomeres. As the cells divide, the telomeric DNA shortens with each successive replication. Soon the telomere becomes too short to maintain its structure as a t-loop and can be accessed by the enzyme. The telomere subsequently re-lengthens and can reform a t-loop. This cycle of shortening and lengthening means the t-loop effectively protects the telomere end.[1]
Maintenance of Telomeres
Telomeres of cancer cells show a remarkable contrast in their behaviour in comparison to those of normal cells. In the right conditions these cells will divide indefinitely and never die, this is why cancer is so difficult to cure. Cancer cells appear to have some form of telomere maintenance pathway capable of invoking this indefinite growth. One of the changes necessary for the immortalization of cancer cells is the ability to bypass senescence therefore the cells will not completely loose their telomeres. Instead their shortened telomeres will be rescued and maintained by adding telomeric repeat units onto the chromosome ends. In this way the genetically disturbed cells gain the immortality characteristic of cancer.
Mechanisms of telomere maintenance
maintenance of telomeres can be undertaken by 2 independent mechanisms;
1. Using telomerase
and2. Using ALT.
The details of these mechanisms are discussed below.1. Using telomerase
This is the most prevalent mechanism, it is based upon the addition of new TTAGGG repeat units to the end of the telomere. This replenishes their DNA and compensates for the telomere shortening that would accompany cell division. In a cell where telomerase is present, the overall length of telomeres is regulated within defined boundaries. By enabling the cells to maintain a stable telomere length, the chromosome end segments are protected. This provides the opportunity for the sustained proliferation that characterizes cancer.
Not surprisingly a special enzyme called telomerase, appears to exist solely for the purpose of maintaining the telomeres. In human cells telomerase is only found to be active in sperm, ovum and during the development of the embryo. In these situations it is necessary for the cells to repair their telomeres. Once the body is fully formed and telomeres of sufficient length have been synthesised, the telomerase activity is shut off in the majority of cells and their telomeres begin to shorten as they reproduce. In germline cells and some other cells involved in the immune system, telomerase remains active in order to keep the length of their telomeres constant.[15]
One of the fundamental features that distinguishes a cancer cell from a normal cell is its ability to divide indefinitely, it is therefore not surprising to learn that an identical situation is apparent in cancer cells. Telomerase activity has been detected in around 85% of malignant tumours tested so far.[17] This correlation suggests that a common mechanism for telomere maintenance in cancer cells is through the reactivation of telomerase in these cells. Since telomerase has been found to exist in numerous types of cancers, inhibition of this enzyme could be a potential cure for many of these.
Further more it is important to note that normal cells that are adjacent to the tumour have shown low or undetectable levels of telomerase activity.[16]
What is telomerase?
Telomerase is a naturally occurring enzyme, that catalyses the extension of telomeres in our cells. This means that telomerase is able to speed up the rate of this reaction but does not get used up itself. Telomerase actually comprises of two components, both of which are essential for telomerase to be active in a cell.
The first component of telomerase is called hTERT, it is a single-stranded overhang of DNA at the end of the telomere. hTERT catalyses the lengthening of the telomere by acting as a template for the second telomerase component to bind to.
Fig 11. The Mechanism of Telomerase
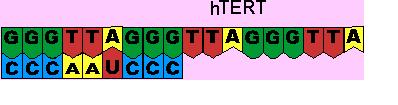
The second component of telomerase is called hTR, which acts as a template for replication.[18] hTR is approximately 445 nucleotides long. Within these 445 nucleotides, 11 nucleotides can be found with a unique sequence of bases, CAAUCCCAUC. This sequence acts as a binding site for the nucleotide sequence found at the ends of telomeres, TTAGGG.[19,20] Initially the binding site on hTR, recognizes the end of the telomere. This involves base pairing interactions between the A-U and G-C nucleotides of the hTERT and hTR components.
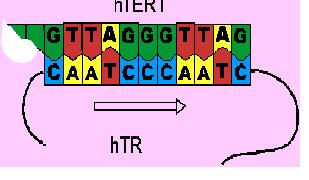
After the initial recognition of the telomere end, the binding site of hTR interacts with the telomere end in such a way that it can be used as a template. It guides the addition of the six-base telomeric repeat unit, GGATTG, to its telomere end. This is called elongation.
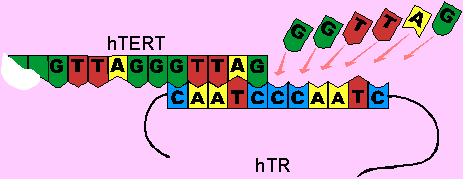
The hTR then moves along the hTERT without dissociation, thus allowing another repeat unit to be added. This is called translocation.
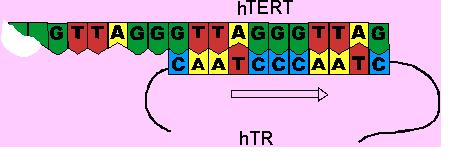
By adding to the ends of telomeres, telomerase ensures the net maintenance of telomere length. By lengthening the strand prior to its replication, it thus becomes possible to compensate for the telomere shortening which follows during the replication of the DNA.
2. ALT
Although in most cases telomere maintenance in human cells is linked to the presence of telomerase, telomerase is not activated in a minority of tumours, around 15%. Some cells compensate for the absence of telomerase and their telomeres can sometimes be sufficiently maintained to permit indefinite proliferation of cell populations. This telomerase independent mechanism has been termed ALT, standing for Alternative Lengthening of Telomeres. In these cases progressive shortening of telomeres was not observed. Instead the telomeres tended to be longer than in the telomerase-positive cells and also showed an increase in length after immortalization.[21]
The mechanism appears to involve recombination processes and has been demonstrated to occur in human cells. Although the mechanism of ALT is not currently known, its existence needs to be considered as a potential resistance pathway to telomerase inhibitors. These ALT mechanisms should therefore be taken into account when considering new methods of inhibiting telomere maintenance.[21]