http://www.ch.ic.ac.uk/rzepa/chimia/index.html#introduction
The first component of this URL up to the # defines a resource, i.e. a mechanism to obtain a document from a designated global location, although it could also be a processing resource such as a search query rather than a document. The string after the # specifies a document fragment, or more generally a definition of actions to be performed on the document or by the processing resource. The old-style session in this model is thus replaced with a collection of hyperlinked documents and resources that together defined a structured set of resources.
MIME and Chemical MIME
Since MIME has been well described elsewhere,2 we will introduce only four commonly used MIME types to illustrate the themes in this article. These are1. text/html 2. image/gif 3. chemical/x-mdl-molfile 4. model/vrmlThe first of these is simply a text-based document, with the expectation that the internal structure of that document will be "marked-up" using the HTML guidelines. Based simply on the globally accepted definition of version 2 of HTML, in combination with equally standard MIME declarations and the URL definition shown above, it had become possible by mid 1994 for a single person, with only limited resources at their disposal, to create a text-searchable index of most of the Internet. Such search facilities are now commonplace, although their utility for retrieving chemical information has been questioned.3 I will return to this theme later in the article.
Finally, we will introduce chemical specifics into this discussion! The very earliest chemical content referenced within HTML type documents was in fact via hyperlinks to images, most popularly defined by a format known as GIF (graphical interchange format) introduced around 1987. Chemical diagrams encoded in GIF (and a few other common formats) were easy to create, and from 1993 to the present, their use proliferated. I argue here that their use as carriers of inter-operable chemical information has been an unmitigated disaster, and we will suffer the problems caused by their continued use for the forseeable future. Why? Well, put simply there is no standard way of identifying that their content is chemical. Only now are computer scientists starting to come up with efficient schemes for scanning the patterns in an image, creating databases of this content, and allowing search and retrieval schemes based on these patterns. The application of image recognition in chemistry remains non-trivial.4 Perhaps a better approach has been to insert chemical information into "hidden fields", which formats such as GIF and PNG (Portable Network Graphics) allow,5 and which would permit a reconstitution of the chemical information by suitable programs. Nevertheless, this approach is rarely adopted, and hence attempts to index the chemical content of the Internet have accordingly encountered enormous difficulties3. An opportunity to insert chemical information in a more simple "human readable" form was missed however. The standard way of invoking an image from a document written in HTML 3.2 or earlier is as follows;
<IMG SRC="sildenafil.gif" alt="C22H30N6O4S" WIDTH="500" HEIGHT="490">
Firstly, note that the actual name of the GIF image suggests chemical content (to a chemist!), but such names are rarely chosen to be unambiguous or complete. The dimensions do not help at all. The so-called ALT field however provides an alternative text-based descriptor of the possible content of the image. In a chemical sense, no guidelines exist for how this field might be used. If present at all, it is normally entered as free-form text; formulae, or even better some form of atom connectivity such as a characteristic SMILES atom-connection string (e.g. O=S(C1=C([H])C([H])=C(OCC)C(C(N2[H])=NC(C(CCC)=NN3C)=C3C2=O)=C1[H])(N4CCN(C)CC4)=O) have rarely been used. The image ALT field however does introduce one important new concept known as meta-data. This is simply a description of an information resource, with the term "meta" deriving from the Greek word for change. Its purpose is to document the origins of, and/or track the change or use of, data.
The HTML 4.0 specification6 replaces the image invocation with the syntax;
<OBJECT title="Sildenafil (Viagra), Molecule-of-the Month at Imperial College" data="sildenafil.gif" type="image/gif" width="500" height="490"> 5-[2-ethoxy-5-(4-methylpiperazin-1-ylsulfonyl)phenyl]-1-methyl-3-n-propyl-1,6-dihydro-7H-pyrazolo[4,3-d]pyrimidin-7-one </OBJECT>
Here, the title is a possible carrier of meta-information, whilst type refers to the media (MIME) type, in some ways another form of meta-data. The remaining text (in this case the chemical name) would only be displayed if for some reason the GIF image could not be shown. This field could serve a dual purpose in providing valuable text-based information as an alternative to attempting to recognise the image content.
Recognising that if chemical content on the Internet was to be rendered indexable, and hence retrievable, we proposed2 in early 1994 that where possible, generic images with chemical content should be replaced by a more explicit declaration. The so-called chemical MIME type was introduced, along with around 20 sub-types that represented a spectrum of chemical content carried by standard or de facto file types that the community had adopted over the preceeding 25 years, and for which tools for their generation and viewing were available to a greater or lesser extent. The molfile format documented comprehensively by MDL is a good example of this MIME type. It can contain either 2D or 3D molecule coordinates, and has an explicit declaration of the atom connectivity and bond types. Put simply, a reference to such a file from within an HTML document carries a strong assumption that an unambiguous declaration of a molecule might be expected. Other chemical MIME types defined other aspects of molecular connectivity, or specified analytical data carried in more or less standard formats which could be generated directly from analytical instruments. The support of this and about 10 other chemical MIME type via browser plug-in software such as Chime from MDL7 or ChemDraw/Chem3D Net plug-in from CambridgeSoft Corporation,8 and via Java-based applets such as ChemSymphony9 and analytical data interpreters10 has ensured that the adoption of chemical MIME has gradually increased from 1994 onwards. A typical example of how this infra-structure could be used to deliver accurate and context-rich molecular information across a range of molecular sciences is shown in Model/Figure 1. This designation is used deliberately; if you are viewing this article in print, then you will inspect a figure (or illustration) of this concept. If you are viewing the article using the Internet, then you can inspect a model.
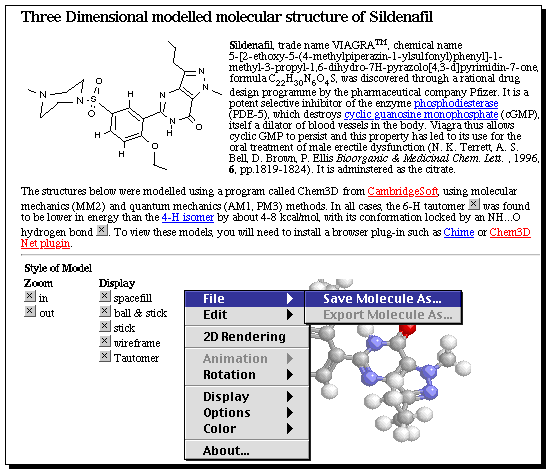
Model 1. Viagra (Sildenafil) as an Internet Model
The model, as opposed to the figure, is invoked within HTML 3.2 as follows;
<embed src=viagra.mol width=300 height=200 name=viagra BGCOLOR="white"
spinx=10 spinz=10 spiny=10 startspin=true options3d=specular display3D=ball&stick
alt="O=S(C1=C([H])C([H])=C(OCC)C(C(N2[H])=NC(C(CCC)=NN3C)=C3C2=O)=C1[H])(N4CCN(C)CC4)=O">
The first line of attributes are generic, whilst the additional forms are specific to the chemical model, and to a large extent, to the software used to resolve this model on the computer screen. The ALT field in this instance is entirely non-standard, but benign in the sense that it is ignored by the modelling software, and would serve only to provide meta-information.
The recommended future method of invoking a model within HTML 4.0 is:
<object data="viagra.mol" width="300" height="200" id="viagra" title="Viagra"
style="spinx: 10; display3D: ball&stick" type="chemical/x-mdl-molfile">
<OBJECT title="Sildenafil (Viagra), C22H30N6O4S" data="sildenafil.gif" type="image/gif" width="500" height="490">
</OBJECT>
5-[2-ethoxy-5-(4-methylpiperazin-1-ylsulfonyl)phenyl]-1-methyl-3-n-propyl-1,6-dihydro-7H-pyrazolo[4,3-d]pyrimidin-7-one
</object>
This defines a cascading order in which attempts will be made by the browser to display either the chemical/x-mdl-molfile, the image/gif or the text field objects. Within the model shown above are also objects which serve to identify small molecular components of the molecule object, such as the region of tautomerism, or a key hydrogen bond, i.e. the objects themselves can have relationships to each other.
Even by 1998, only perhaps a few percent of all molecular content on chemical web pages was identified using MIME types. These chemical MIME types are also largely seen now as a legacy from the days of proprietary formats and programs. Often, these formats lack modern mechanisms for defining internal structures and have to be considered as a single component (a "blob"). One would not expect to easily identify smaller well defined components within the document such as molecular sub-components. Such legacy files also did not provide any definition of a standard mechanism for specifying meta-data. Frequently, it might even prove impossible to identify which version or flavour of file one was dealing with (i.e. tracking the change in use of data with time). Before turning to molecular components and meta-data, one further important MIME type should be discussed.
The Transition from Images to Models
The adoption of chemical MIME types has allowed the chemical content of the Internet to be carried not by the poorly structured figures, schemes, or illustrations, but by what we call the MODEL11 and by much more formally structured document types. Thus a molecule for example can be clearly and unambiguously specified not by an un-indexable image, but by a model, represented for the user's use by appropriate software within a browser window. Most importantly, the chemical model is available for the user to re-use in whatever context they wish. Such chemical models form one basis for what we term the third generation of electronic journals, where the journal is best considered as a tool for placing into context and manipulating the models it delivers.12-14 This contrasts with the much more passive illustrations offered by print, or print-like classical electronic journals.Alongside chemical models, the MIME type model/vrml allows a more generic modelling functionality, known as Virtual Reality Modelling Language or VRML15. This type of model serves to integrate molecular models with complex 3D data representations, molecular surfaces, animations, and most interestingly, processing functionality within the model. Such models can have so-called script nodes associated with model components based on defined algorithms. For example, two molecule components of a VRML model could be associated with a defined force field that could dynamically compute the energy of interaction of the two components during an attempt to dock one model with another. Such composite scenes also allow a much richer integration of chemical with non-chemical models, and these work particularly well at the boundaries between chemistry and other disciplines. Virtual Reality models have been recently reviewed from both a general15 and a chemical perspective.11
In one sense, the problems alluded to above in identifying the chemical content of two-dimensional images are also inherent in three dimensional models. Whilst chemical models generated from formats such as the MDL molfile can be readily indexed and search for, much still needs to be done in chemically indexing the more generic VRML models. The equivalent of the image ALT field or HTML document meta-data could to be found in the so-called VRML Viewpoints, but as with image ALT declarations, no standards in their use are employed, and such viewpoints are not yet indexed routinely by any index and search services. Interesting progress has been made in the re-identification of chemical content from VRML models16 and progress is expected to be rapid in this area in the future.
Meta-data, Molecular Components and Modularity
Meta-data is a description of an information resource, and may be used for a variety of purposes, such as identifying whether the resource meets a particular information need, evaluating the quality or fitness for e.g. defined chemical application of the resource, or tracking the characteristics of a resource for subsequent maintenance or usage over time.A metadata record consists of a small set of attributes, or elements, necessary to describe the resource in question. These include basic attributes of a document such as its title, date, description, creator (author), format (MIME type), subject (keywords) and relation (of the resource to other resources, normally via a URL declaration). The meta-data is normally contained in a particular component of the document known as the header. Its implementation within any particular type of document can vary; that for HTML documents has recently been standardised,17 and proposals for implementation within images and perhaps even VRML types might be expected in the future. One example of using such declaration to enable a resource to be evaluated for a particular need was the inclusion of the following (highly non standard!) meta-data header in each of the articles comprising the ECTOC-3 electronic conference proceedings;18
The so-called chemical prototype attempts to define a single molecule that best represents the overall molecular content of document. In principle, this would allow automated analysis of the document to provide an indication to a user of whether the chemical content is close to their interests, or conversely whether the document represents a "dissimilar" contribution in a scan of molecular diversity. Because agreement on meta-data types and syntax within HTML documents has only recently been formalised as the Dublin Core standard,17 the five year evolution of the Web has assimilated few of these guidelines. Almost no HTML documents contain any significant meta-data declarations, and of those that do, even less are chemically useful. As an example, the on-line version of this article contains some Dublin Core declarations. A project to define a discreet set of standard chemical meta-data declarations such as coordinates, substance, computation/simulation, biological activity, safety, synthesis, characterisation, instrumentation, physicochemical data, reaction data and crystallography (provisionally christened Dublin Chem) is under way.19
The chemical document illustrated in Figure/Model 1 above illustrated how discrete molecular components could be identified within a larger molecular model. This is a very specific example of a general problem in chemistry as a whole. The underlying documents used to create Model 1 utilised a combination of HTML to define the text and links between objects, MDL molfiles to carry small molecule connectivity and 3D coordinates, CSML (chemical structure markup language) to define molecular fragments and Brookhaven PDB files to carry macromolecular information. In the summer of 1995, a project was started20 to define a chemical equivalent of HTML that would serve to provide a single self-consistent syntactic framework to replace these ad hoc methods with modular components. The latest version of CML (Chemical markup language) follows a set of guidelines known as XML (extensible markup langauge)21 and is described in detail in another article in this issue.22
Conclusions
Prior to the introduction of Internet-based document delivery systems such as the World-Wide Web, a significant proportion of on-line chemical information was held in the form of proprietary databases, almost invariably requiring user authentication and custom software for access, but also offering high and consistent quality. Since 1993, around 2 million Internet-accessible documents have been indexed based on their title or the appearance of the words chemistry or chemical in the text. A significant proportion of these might be expected to contain links to images with further, but undetectable chemical content, and the overall quality of these documents is highly variable. The number of documents with strongly typed molecular information, i.e. apparently around 7200 files in PDB format, 1400 MDL molfiles, 1200 XYZ, and 130 J-CAMP spectral files, comprise only around 0.5% of the total supposed chemical documents. Of these 2 million chemical documents, it is supposed that few carry meta-information, alternative chemical fields or links to models, a supposition because the current generation of search engines are not programmed to search for such information. There is no doubt that the quality of the retrievable chemical information on the Internet would rise substantially if more use was made of the types of mechanisms described above.Another approach to creating chemical documents involve generating them dynamically from closed databases in response to a suitably constituted request or search query. For example, the ChemFinder site3 contains a large amount of chemical information, but this presented to the user only in the form of a "just-in-time" document, and hence this content is not reflected in the statistics quoted above. It may well be that the majority of useful Internet-based chemical information will become available only via such controlled molecule or journal databases, created by a large number of authors, but controlled by a small number of publishers, a situation which reflects of course the current situation in printed publishing. The Internet does offer an alternative paradigm, in promoting the use of chemical models and other high value and modular chemical data in an open manner, and one where entirely new "added value" models of chemical resource discovery and "Knowledge Capture for Compounds" could be created using information management techniques. If this second scenario is to come about, then the creators of this information will have to make it happen. If we adopt the same models that lead to the creation of an over-abundance of paper-based information graveyards, then it will probably not happen. The future is in our hands.
References
- H. S. Rzepa, Science Progress, 1996, 79, 97; H. S. Rzepa, P. Murray-Rust and B. J. Whitaker, Chem. Soc. Revs, 1997, 1-10.
- P. Murray-Rust, H. S. Rzepa, B. Whitaker, J. Chem. Inf. Comp. Sci., 1998, September issue.
- J. Brecher, Chimia, 1998, 52, this issue; W.-D. Ihlenfeldt, ibid. See also S. J. Clarke and P. Willett, ASLIB Proceedings, 1997, 49, pp.184-189
- A. Simon, A. P. Johnson, J. Chem. Inf. Comp. Sci., 1997, 37, pp.109-116.
- For further details, see W. D. Ihlenfeldt; http://www2.ccc.uni-erlangen.de/services/gif.html
- D. Raggett, A. Le Hors and I. Jacobs; http://www.w3.org/TR/REC-html40/
- B. A. Van Vliet, T. M. Maffett, Abs. Papers Amer. Chem. Soc, 1997, 214, pp.177-COMP. See also http://www.mdli.com/chemscape/chime/
- A. Hinchliffe, Elec. J. Of Theor. Chem, 1997, 2, 215-217. See also http://www.camsoft.com/plugins/chem3D.html
- P. Tebbutt, A. Hodgkin, A. Krassavine,Abs. Papers Amer. Chem. Soc, 1997, 214, pp.58-CINF. See also http://www.cherwell.com/chemsymphony/
- Viewers are available for the JCAMP standard; A. N. Davies, Analyst, 1994, 19, pp.539-542, http://www.isas-dortmund.de/projects/jcamp/, and the SPC analytical chemistry format (http://www.galactic.com/). See for example a viewer available as a Java applet; http://www.ch.ic.ac.uk/java/applets/jspec/spectra/
- O. Casher, C. Leach, C. S. Page and H. S. Rzepa, Chem. Brit., 1998, in press. For earlier articles, see O. Casher, G. Chandramohan, M. Hargreaves, C. Leach, P. Murray-Rust, R. Sayle, H. S. Rzepa and B. J. Whitaker, J. Chem. Soc., Perkin Trans 2, 1995, 7; C. Conesa and H. S. Rzepa, Perkin Trans 2, 1998, 857.
- CLIC Project: D. James, B. J. Whitaker, C. Hildyard, H. S. Rzepa, O. Casher, J. M. Goodman, D. Riddick and P. Murray-Rust, New. Rev. Information Networking, 1996, 61; H. S. Rzepa, B. J. Whitaker, J. Goodman, O. Casher, D. Riddick, J. Griffiths, D. James, C. Hildyard, Abs. Papers Am. Chem. Soc, 1997, 214, pp.45-COMP. For examples of articles enhanced as part of the CLIC project, see http://www.rsc.org/is/journals/current/chemcomm/cccenha.htm
- Journal of Molecular Modelling: T. R. Clark, Advanced Materials, 1996, 8, p.787; H. H. K. Roth, T. R. Clark, Abs. Papers Amer. Chem. Soc, 1997, 214, pp.43-COMP.
- Internet Journal of Chemistry: S. M. Bachrach, D. C. Burleigh and A. Krassavine, Issues in Sci. Tech. Librarianship, 1998, No. 17.
- W. D. Ihlenfeldt, J. Mol. Modeling, 1997, 3, pp.386-402.
- M. Leipold and W. D. Ihlenfeldt, personal communication: http://vermeer.organik.uni-erlangen.de/dissertationen/
- S. Weibel, Bull. Amer. Soc. Inf. Sci, 1997, 24, pp.9-11. For a user-guide, see http://128.253.70.110/DC5/UserGuide4.html
- ECTOC-3: (Eds H. S. Rzepa and C. Leach), Royal Society of Chemistry, 1998, ISBN (CD-ROM) 0-85404-889-8.
- P. Murray-Rust and H. S. Rzepa, to be published.
- The CML project was first described in P. Murray-Rust, C. Leach and H. S. Rzepa, Abs. Papers Am. Chem. Soc , 1995, 210, pp.40-COMP; P. Murray-Rust and H. S. Rzepa, Abs. Papers Am. Chem. Soc, 1997, 214, pp.23-COMP.
- P. Murray-Rust, "Chemical Markup Language, A simple introduction to structured documents", in "XML, Principles, Tools and Techniques" (Ed. D. Connolly,), O'Reilly, 1997, pp 135-149.
- P. Murray-Rust, Chimia, 1998, 52, this issue.