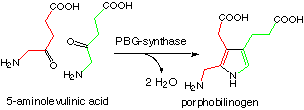
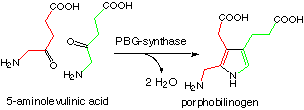
PBG-synthase catalyses the asymmetrical condensation of two 5-aminolevulinic acids to produce PBG.
We are working on the PBG-synthase from E. coli CR 261. This is an enzyme with a molecular weight of 280 kD constituted of eight identical subunits. The eight subunits are postulated to be organized as four functional units each composed of a dimer of the protein. The PBGS from E. coli is Zn2+ and Mg2+ dependant and shows an optimal pH value around 8.
The active site recognizes the two substrate molecules but shows a greater affinity for the first one which binds to the enzyme covalently. The amino function of a lysine side chain reacts with the keto function of the 5-aminolevulinic acid (ALA) to form an E-iminium ion [1]. Jordan [2] located this first substrate in the P-side whereas Shemin [3] postulated that the first substrate is the A-side ALA.
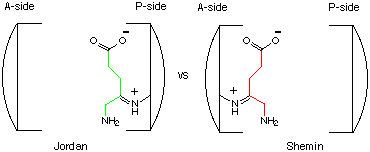
The second substrate seems not be bound covalently to the enzyme. However the keto function is also essential for a good recognition of the second substrate. The carboxylate function is important for the recognition of the A-side ALA whereas the amino group has been shown to be less important.
A complex can be postulated between the second substrate and the ZnA2+ [4] which is essential to the enzyme activity.
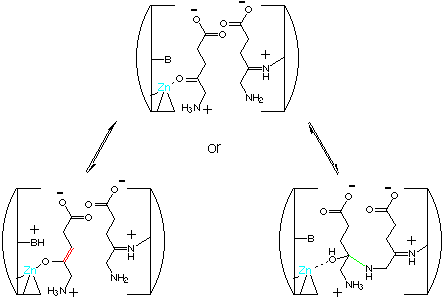
Our kinetic model is based on Michaelis-Menten kinetics with a rapid formation of the Schiff base between the enzyme and the first substrate at the P-site :
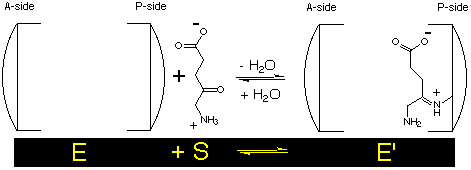
The next step is the complexation of the second substrate at the A-site and then followed by the formation of the product which is the rate determining step :
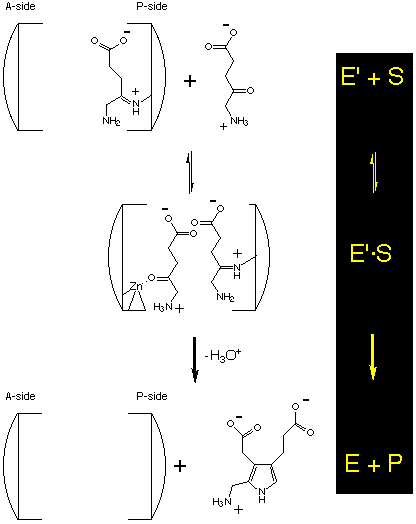
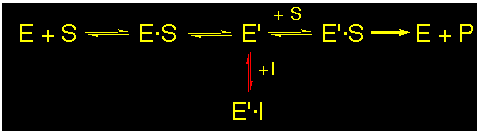
Uncompetitive or mixed :
The inhibitors interact with the P-site (uncompetitive) or with the P-site and the A-site (mixed) of the active site of the enzyme at the same time. Mixed or uncompetitive inhibitions can not be clearly differentiated by the interpretation of kinetic data only.
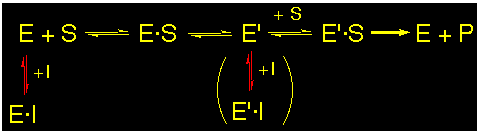
Slow-binding :
The slow binding inhibitors show a time dependent reversible inhibition. The inhibitors form a sufficiently well stabilized interaction with the enzyme (e.g. stabilized Schiff base at the P-site of the enzyme) to dissociate only slowly under dialysis conditions.
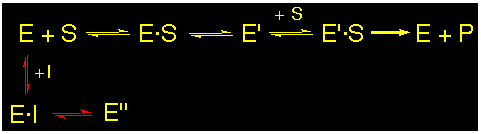
Competitive :
Inhibitors belonging to this class interact with the A-site of the enzyme.
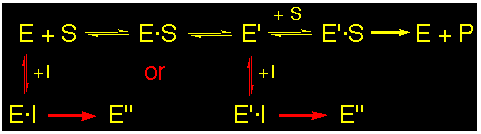
Irreversible :
These inhibitors bind to the enzyme covalently at the active site either at the P-site or at the A-site of the enzyme.
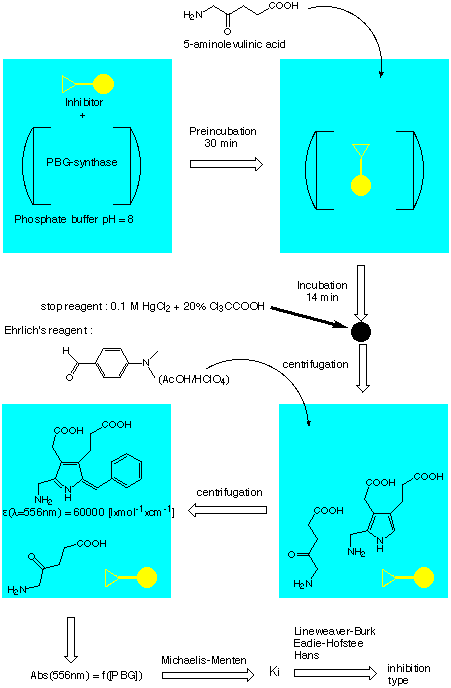
The procedure for testing the inhibitors is schematically shown bellow :
4,6-dioxoheptanoic acid (1) is known to be responsible for the inhibition of PBG-synthase in the disease called tyrosinemia. Tyrosinemia has its name due to accumulation of 4,6-dioxoheptanoic acid (1) as a product of the abnormal degradation of tyrosine [5]. Testing of 1 as inhibitor with the enzyme from E. coli. Gave the smallest inhibition constant determined so far. The Ki-value of 1.4 µM is almost a factor of 10 smaller than inhibition constants determined for the next best inhibitor. The inhibition kinetics was typical for an uncompetitive or mixed inhibitor. We interpret this behavior as a consequence of an interaction of the inhibitor 1 at the P-site and may be also at the same time at the A-site of the enzyme.
The goal of our investigation is to assess the impact of structural modifications of the 4,6-dioxoheptanoic acid (1) on the efficiency as an inhibitor of PBGS. We hope to contribute to the understanding of the role of the 1,3-diketo system in the interaction with the A-site and with the P-site of the enzyme.