Henry S. Rzepa
Abstract: A review of the development of computational chemistry tools which make direct use of the Internet is presented, together with some recent advances in new Internet methodologies based around Java Applets and VRML (Virtual Reality Modelling Language).
Keywords: Internet, World-Wide Web, VRML, Java.
___________________________________________
Internet Based Computational Chemistry Tools: 1970 - 1992.
The use of computers as tools in chemistry dates back to the late 1950s and the adoption of Fortran as a scientific programming language. Much of that history relates to data entry, its transfer and processing via suitable programs, and display of the processed results on a suitable local device. The network connecting these various devices during the period 1970 - 1980 invariably comprised a simple point-to-point connection between devices. Only from around 1980 - 1990 did these networks develop into a switched model or inter-networks, where the distinction between the computer and the display devices vanished and where the user had access to a variety of network based resources from a single point of use. Around this time, a particular collection of protocols defining the movement of data became associated with the generic term "Internet". Specific implementations of these TCP/IP standards such as the Telnet terminal protocol and the FTP file transfer protocol became well known, and widely used to relocate scientific programs and information around the globe. For the first time a global infra-structure was introduced down to the desktop level for interconnecting computers.
Up to about 1992 however, the Internet was almost exclusively used as a carrier for proprietary programs and information, encoded in such a way that there was very little inter-operability for the chemical information that was exchanged. For example, the Telnet terminal interface functioned as a 80 column and 24 line display of ASCII text characters. Its implementation for personal computers around 1988 enabled copy-and-paste operations to be performed on this text, but as a computational chemistry tool, this was really a very primitive medium. Some file formats for example depended on line-breaks to encapsulate the semantic content, and these could not always be relied on to be transmitted correctly since the three major operating systems in use (Unix, Windows and Macintosh) encode these line breaks differently. Graphical display was handled during the 1980s by the Tektronix vector graphics format. Here, the ability to "copy" a structure from a graphical display window and "paste" live 2D or 3D coordinates into say a structure editing program was almost entirely absent. Molecular information rarely arrived in a form which enabled its re-use outside the proprietary programs it was designed for. In effect then, the Internet functioned as a transport device for a variety of point-to-point interchanges of information, and could not then be considered as an integrated medium for chemical manipulation.
In 1992, the first serious attempt at integration became widely adopted. The Gopher system[1] was a client-server model comprising a smoothly integrated hierarchical file system operating on a global scale, rather than being limited to single computers or small clusters of computers. Clients for using the Gopher system became widely available for the majority of computer platforms, and the system also had a functioning global search facility known as Veronica. In 1993, Gopher+ introduced a feature known as MIME (Multipurpose Internet Mail extensions). MIME had been developed as a mechanism for attaching documents to mail messages, and comprised a standard way of placing a header (and if necessary delimiter) which would uniquely identify the form and structure of the attached data. Its use with the Gopher system enabled for the first time the interchange of other media types such as images, animations and sound. Our own first use of this system in 1993 was associated with the archiving of supplemental information in conjunction with research papers published in printed journals.[2] Thus diagrams which made extensive use of colour could be made available at no cost to readers of the articles. We also investigated the use of video animation to illustrate the three dimensional properties of computed molecular surfaces such as electrostatic potentials.
Although much more of a genuine Internet based computational chemistry tool, Gopher suffered from several defects.
i. Text, images and animations had to be kept as separate files in the file-system, and the user had to make a separate request for each. Text was displayed using the Gopher client, but the user had to supply further programs ("helpers" as they became known, for processing other types of file).
ii. The information as received by the user was still largely not inter-operable. An image or animation could really only be interpreted by a human mind, and its re-suability from there was still subject to human error!
iii. There was no real interface into existing "legacy" resources such as databases, library resources, computational resources etc.
iv. No mechanism existed for background reprocessing of information using defined algorithms and hence presentation to the user in a ready-to-use form.
v. There was no widely adopted mechanism for defining how user supplied information could be remotely processed and subsequently returned to the user.
The Period 1993-1995.
In late 1993, two major developments came to the fore which addressed most of these limitations. The World-Wide Web[3] added a new language known as HTML (Hypertext-markup-language) and a new Internet protocol for transferring this information (HTTP). HTML introduced many concepts which had originated from the publishing industry in the form of SGML (standard generalised markup language), where the central idea was of precisely structured information which could be subjected to exact parsing against a description of the document structure known as a "document type definition" or dtd. Effectively, HTML took many concepts that had been under development in the publishing area since the middle 1980s, and integrated them with the Internet communications infra-structure via a device known as a URL (Uniform Resource Locator). Such a URL specified the precise location of a "resource" such as a document, a search engine, a computational chemistry database or supercomputing centre, along with the exact protocol for how this resource should be returned to the user. It is the far-sighted use of the word "resource" that reinforces the claim that a new computational chemistry tool had been created. The URL was integrated into an HTML based document using a device known as a hyperlink, or more precisely an "anchor". This coming together of the ideas of structured (and therefore inter-operable) information coupled with a unique global addressing system for this information represents a key defining moment when the Internet first achieved its distinct status as a computational chemistry tool rather than simply being a transportation medium.
Also in 1994, we recognised[4] that there were effectively no standards in place which recognised the unique character of chemical information (as opposed to text, audio, video etc). We introduced a draft paper[5] which proposed a set of standard chemical media types to be used with the MIME protocols noted above. What did this combination of the World-Wide Web, the HTML language, the URL and chemical MIME allow one to achieve?
i. It allowed authors to create a new form of document, one in which a descriptive narrative of a chemical discovery could be integrated into "live" chemical information and resources. The data which the authors had acquired and which they had interpreted, could be made available to the readers of the information. Most importantly, this data need not reside on a single computer system, but could be choreographed from a variety of sources.
ii. This system also achieved a degree of separation of the chemical "content" from the chemical "style" or "form". No longer was the reader of the information bound to accept the particular visual style or interpretation of the data imposed on it by the author. Instead, if they wished, they could ask that the data be passed to their own visual interpreter or indeed much more sophisticated form of processing tool. Perhaps a specific example will emphasise this point. Up to around 1993, 3D molecular coordinates, whether obtained from experimental or computational studies, had to be reduced to a single 2D projection, often with any colour eliminated, for printing in a conventional journal. That is still very much the case with the journal you are currently reading, where pages charges for colour reproduction are very high. The reader of such a journal, if they really wanted to re-interpret this data, would have to wait perhaps two years for the coordinates to become available via a suitable database. Even then, they would have to license the data base and perhaps the programs needed to search it, before they could gain access to this information. Now, the World-Wide Web allows the 3D coordinates to be integrated into the document describing the research. The client program that is used to connect to the World-Wide Web (typically Netscape or Internet Explorer) can be enhanced with a chemical MIME aware "plug-in" such as Chime.[6] This combination allows the user to acquire a chemical document from the Internet, and to impose a molecular style on the content such that any 3D molecular coordinates are integrated directly into the screen display. Thus where text is used to describe molecular behaviour, the user can cut-and-paste this into whatever program they might find useful for reprocessing the information. If they see a molecule in the same document, not only can they rotate the 3D representation of the molecule, change its stylistic attributes from e.g. wirefame to e.g. spacefill, decide which colours to display it in, perhaps measure interatomic distance, even perform computations on the system, but most importantly, they can save all this information either to hard disk, or perform the equivalent of a copy-paste operation from the Web client window to some other program of their own choosing.
Perhaps the earliest example of how such "hyperactive" molecules were seamlessly integrated into a chemical document is in a keynote paper for the ECTOC electronic conference.[7] The concept has also been extensively used in the later ECHET96 conference[8] and in an electronic version of the Journal "Chemical Communications"[9] for enhancing the keynote articles. In summary, the World-Wide Web, in conjunction with chemical MIME headers, turns a chemical document into a working live tool, where the user makes the decisions on what to do with the data, rather than simply accepting a single point of view imposed by the author of the information.
iii. Two other important concepts were introduced at this time. The first was the ability of the user to provide their own parameters via a built in "form" interface. in the HTML document. These parameters could be transmitted to a remote server, and acted upon by programs and scripts via a second generic concept known as "cgi" (common gateway interface). This allowed entry to the vast "legacy" of databases, and functionality already existing on the Internet. The results of whatever action the scripts and programs performed could be returned to the user via the Web client window.
These three themes were illustrated "live" in the oral presentation corresponding to this article, given at the WATOC96 conference in Jerusalem in July 1996.[10] In converting this talk into a version capable of being printed in monochrome or grey scale tones, some interesting and largely non-scientific issues come to the fore. Firstly of course, there is little point in projecting the 3D images shown into a 2D plate for reproduction in this journal, since this represents not only a loss of information, but a reduction in the usefulness of the "tool". Secondly, although the publisher of this journal does maintain a Web site for precisely this purpose in order for supplemental material to be mounted there,[11] the author currently has to transfer copyright to the publisher. Copyright law in this context relates to text and images. Little precedent exists for how to handle live "working" tools which happen to be an integral part of a scientific publication of the type that are discussed below. For this reason, the reader is referred not to the publishers Web site, but the author's own Web site for a "working" illustration of this article.
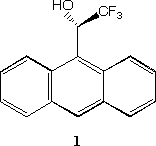
Three problems in chiral recognition were presented to the audience.[12] The chiral properties of these molecules are intrinsically bound to their 3D molecular structures. These structures were shown to the audience with the important aspects of each molecule such as the chiral centres rotated into view in each case, or any hidden aspects revealed. Furthermore, via a simple scripting extension, the key centres could be highlighted and the attributes of these centres changed to e.g. spacefilling representation. In the scenario presented, it became clear that further properties of e.g. molecule 1 should be computed, namely in our case the molecular electrostatic potential of the [pi]-electron region.

Its at this point, that the immaturity of the Internet as a computational chemistry tool starts to be manifest. In order to edit the 3D molecular coordinates of molecule 1 into a form suitable for computation, they would have to be transformed from Brookhaven PDB format to e.g. Gaussian input, and the suitable keywords, a basis set etc added. The program used to visualise the 3D coordinates (Chime) is not really suitable for this, and the user would have to resort to custom programs that they already possess for this task. Thus we come up against two undesirable features of the current Internet.
i. The user is still faced with having to install on their local computer system a variety of "helper" programs or plug-ins etc to perform various specific tasks on the live molecular information they acquire. Even worse, even if the user does install a set of such helpers on their system, there is no guarantee that these programs will inter-operate. For example, the Chime plug-in used to render PDB files cannot be used to "communicate" with the Gaussian ab initio program in the context that we require here.
ii. Many of the existing formats defined in the chemical MIME standards are far from having the structured content as prescribed by the SGML guidelines. HTML itself has essentially no syntax for molecular content (superscripts and subscripts for formulae are about the best it can offer) and as a result one is always faced with the Babel like problem of converting from one format (e.g. PDB) into another (e.g. Gaussian input), with the invariable loss of information this normally results in, and with no mechanism for easily providing the occasional extra information that might be needed (e.g. the spin state of the molecule is not carried in PDB files, but is needed in Gaussian).
True Internet Computational Chemistry Tools.
In 1996, two solutions to these problems were beginning to emerge. The first attempts to construct an infra-structure for inter-operable applications in a manner which is extensible, does not lead to proprietary turn-key solutions into which the user is locked, and which solves the problem of how to deliver the functionality to the user without repeated installation tasks. Based on an object oriented language known as Java (itself similar in construction to languages such as C++), it uses the concept of Class libraries to define inter-operable molecular functionality. The most important aspect of Java however is that the Internet is an intrinsic aspect of its functionality. Firstly, the Java language is used to write small code units known as "Applets", which can be delivered to the user in the same way that a document is delivered. In this sense, Java introduces what has been called "just-in-time" applications, in which the user does not have to pre-install any application on their local system, but acquires each applet only when they need it. Applets themselves, because they derive from object oriented constructions, can themselves call any object class them may need via a suitable request to the Internet, or to a class library that may already be present on the users own local disk system. Again, because of the intrinsic object orientation, potentially any Java Applet can interchange data with other applets, i.e. the problem noted above with the Chime plug-in and the Gaussian program is at least potentially eliminated.
In 1996 of course, we are quite some way away from having e.g. a complete ab initio program functionality present as a Java molecular class library, but there is no reason why this could not be achieved in the future. For example, if we had requested simply a B3LYP/6-31G(d,p) calculation of the molecular electrostatic potential on a molecule containing only C, H, O and F atoms, only those Gaussian modules and basis functions needed for this calculation have to be acquired. Another feature of the Java environment (it is more than just a programming language) is its built-in platform independence. Conventional applications are compiled with machine specific compilers to produced executable instructions that only run on a very specific computer and operating system. For example, the Chime PDB viewer is available for SGI Unix workstations, but not for other Unix implementations. Java applets are transmitted in what is called "byte code" form, which are interpreted by what is called a "virtual machine" on the user's system. Such a virtual machine can be implemented within a World Wide Web browser, and thus the availability of the applet is directly related to the availablity of a suitable Web browser. Virtual machines come with one significant limitation; the speed with which they interpret and hence execute the Java applets. In 1997, this concept will be augmented by so-called "just-in-time compilers", also implementable in e.g. the Web browser, which should address this problem.
In 1996, a number of molecular Java applets, including editors, visualisers, sequence alignment editors and a Gaussian front end, had become available from various sources.[14] In order bring some standards to this development process, to document the subject in a manner suitable for molecular scientists and to try to ensure future inter-operability is achieved in the area of molecular science, an organisation known as the "Open Molecule Foundation" has been set up to facilities developments in this area, and to provide information and support for developers and users.[15]
The second major theme in 1996 was to address the problem of legacy and relatively unstructured chemical information formats. The original PDB format form example was characterised by a 80 column format which derived from the days of punched cards, where the marker separating individual atoms or residues is the line break. Over the years, the PDB format has acquired many flavours and enhancements which can results in significant ambiguity in its interpretation. Recognising the need for a structured chemical data format that includes the important principles derived from a modern set of guidelines such as SGML such as separation of content and style or form, the "chemical markup language" or CML has been implemented by Murray-Rust.[16] This functions as a medium for the inter-operability of chemical information between areas such as publications, programs, equipment, databases, other structured file formats such as CIF, CEX, asn.1 and CXF, and older legacy formats such as PDB. Because it is derived from a formally defined SGML "dtd", it can be parsed using standard tools, and because of the highly structured nature of its implementation, it can be associated with the molecular class libraries defined by Java in a highly natural way. Thus the combination of Java technology and CML chemical semantics provides a powerful and highly extensible way forward for the development of Internet based chemical tools.
One area where in 1996 Java was severely limited was the implementation of efficient 3D molecular class libraries, severely restricting it as a 3D visualising and rendering tool. This is an area of course where computational chemistry tools are traditionally well developed. There may be no need however to re-invent the wheel in this category. Consider first however the traditional approach to 3D rendering, exemplified perhaps by codes such as RasMol and its implementation as a Web browser plug-in in the form of Chime. Chime is essentially a black box for converting 3D molecular coordinates to a pleasing visual form. However, such a product does not allow other forms of visual representation or property to be rendered; for example the molecular electrostatic potential calculated for 1 at the B3LYP/6-31G* level. In particular, we wished to show the audience that the negative electrostatic potential was highly chiral, a property that we hypothesis is related to the excellence of this reagent as a chiral resolving agent via its weak binding properties. Such 3D rendered properties still require extensive machine resources for their display, and writing efficient code to do this is still difficult and expensive.
In 1995 a generic solution to such problems was introduced with a language called VRML (virtual reality modelling language). It is an object oriented 3D scene descriptor, much in the way that HTML is a 2D descriptor where the objects are the standard ASCII character set. Because VRML can display 3D objects such as lines, spheres, polygons etc, it is ideally suited for the problems of 3D chemical object and property display. The ability to display VRML objects is now integrated into Web browsers such as Netscape, and hence the problem of writing rendering code is moved from a specific chemical problem to a generic browser issue. In 1996, VRML 2.0 was introduced which offered greater flexibility in defining various 3D objects, and importantly introduced a mechanism for integrating "actions" on the individual objects in a 3D scene using scripts and algorithms based on the Java concept discussed above. In this scenario, Java is used to act upon underlying molecular data, and VRML is used to convert this information to a form which can be presented to the user on the screen. Because VRML is a 3D format, the user can rotate, and navigate this information on the screen under their own control. We have previously shown how[17] VRML can be used to present a variety of types of molecular information to the user in an integrated manner. In the examples presented in the WATOC96 talk, ball-and-stick structure diagrams were overlaid with molecular electrostatic potential maps, computed Connolly surfaces, and 3D scatter diagrams derived from information retrieved from the Cambridge crystallographic database.[18] Just as with HTML, VRML objects can contain URL hyperlinks to other Internet based resources. In our case, individual points in the 3D scatter diagrams could be hyperlinked to other VRML scenes containing details of inter-molecular interactions relevant to the chemistry being discussed, and these in turn can be hyperlinked to bibliographic information about each molecule, or electronic journal articles etc. Other possibilities include linking VRML objects to Java applets or scripts which can perform actions on such objects. The most trivial example would be to change the radius of a spacefill display of any particular atom from e.g. the van der Waals value to some other. A more sophisticated example is the use of Java to display digital spectral information derived from an NMR spectrometer,[19] and to link regions of the spectrum to specific atoms or residues in a 3D molecular object displayed using e.g. VRML. The links can be bi-directional, i.e. clicking on a specified atom will highlight the spectral region containing peaks associated with that atom.
The potential of this combination of Java for interpreting structured chemical information encoded in formats such as CML, and the use of VRML for its 3D display seems immense, and we are currently only at the start of the development curve. Certainly the path forward for developing a new range of computational chemistry tools in which the Internet is an intrinsic part of their functionality has been laid. Many exciting challenges in applying and using these tools await us. These include their application to electronic journals, conferences, books, databases and teaching aids. Issues such as security and validation, the implementation of cost recovery and charging models, how intellectual property rights and priority claims are handled, and how archival mechanisms can be developed in an areas where the technological advances of a few months can easily obsolete formats, need to be resolved. There can be little doubt however that after about 25 years of development, the Internet must now be considered as a major tool as the disposal of molecular science. Indeed, two of our projects the "3D Virtual Chemistry Library"[20] and the "3D Virtual Chemistry Laboratory"[21] strongly reflect this new found role for the Internet.
Acknowledgements: The author thanks his colleagues Omer Casher, Christopher Leach, Christopher Page, Syvain Comiti, Peter Murray-Rust, Jurgen Brickmann and Adam Precious for their inspiration and collaboration on many of the projects cited here and for their continuing enthusiasm in this new cause. Funding from the JISC Electronic libraries (e-Lib) and JTAP programs, from British Telecom for a University Development award and from GlaxoWellcome is gratefully acknowledged.
[1] S. M. Bachrach, "Chemistry and Gopher", chapter in "The Internet: A Guide for Chemists", Ed. S. Bachrach, American Chemical Society, 1995.
[2] See gopher://chcmga.ch.ic.ac.uk/11/Scientific_publications/rzepa/ Royal_Society_of_Chemistry/Chemical_Communications/3_02351F as an archive for O. Casher, D. O'Hagan, C. A. Rosenkranz, H. S. Rzepa, N. A. Zaidi, J. Chem. Soc., Chem. Commun. , 1993, 1337.
[3] H. S. Rzepa, "Chemistry and the World-Wide-Web", chapter in "The Internet: A Guide for Chemists", Ed. S. Bachrach, American Chemical Society, 1995.
[4] O. Casher, G. Chandramohan, M. Hargreaves, C. Leach, P. Murray-Rust, R. Sayle, H. S. Rzepa and B. J. Whitaker, J. Chem. Soc., Perkin Trans 2, 1995, 7; H. S. Rzepa, B. J. Whitaker and M. J. Winter, J. Chem. Soc., Chem. Commun., 1994, 1907.
[5] For general information about this topic, see http://www.ch.ic.ac.uk/chemime/
[6] See the URL http://www.mdli.com/ Chime is itself a Netscape plug-in enhancement of the RasMol molecular viewer written by Roger Sayle. For the history of RasMol, see http://www.glaxowellcome.co.uk/netscape/software/history.html
[7] A. Padwa, E. A. Curtis, V. P. Sandanayaka, and M. Weingarten, ECTOC, 1995, See http://www.ch.ic.ac.uk/ectoc/papers/01/
[8] See http://www.ch.ic.ac.uk/ectoc/echet96/
[9] The CLIC Project. See http://chemcomm.clic.ac.uk/
[10] The on-line version of which is available the the URL: http://www.ch.ic.ac.uk/rzepa/watoc96/
[11] See http://www.elsevier.nl:80/section/chemical/theochem/menu.htm
[12] For further details, see D. O'Hagan and H. S. Rzepa, Chem. Commun, 1996, in press. An electronic version of this article will also be available via the journal home page; http://chemcomm.clic.ac.uk/
[13] For the live example, see http://www.emsl.pnl.gov:2080/forms/basisform.html
[14] For one such collection, see http://www.ch.ic.ac.uk/java/
[15] For information about Java and tools for Bio and Chemo-informatics, see the Open Molecule Foundation; http://www.ch.ic.ac.uk/OMF/
[16] P. Murray-Rust; http://www.dl.ac.uk/CBMT/cml/cml06f/newintro/role.html
[17] For chemically oriented examples of how VRML has been applied, see O. Casher and H. S. Rzepa, J. Mol. Graphics,1995, 13, 268; H. Vollhard, C. Henn, G. Moeckel, M. Teschner, J. Brickmann J. Mol. Graphics, 1995, 13, 368; J. Brickmann, H. Vollhardt, Trends In Biotechnology, 1996, 14, 167-172.
[18] O. Casher, C.Leach, C. S. Page and H. S. Rzepa, Theochem, 1996, September issue;
[19] H. S. Rzepa, P. Murray-Rust and R.Kinder; See http://www.ch.ic.ac.uk/java/HyperSpec/
[20] See W. Locke and H. S. Rzepa, http://www.ch.ic.ac.uk/vchemlib/
[21] See A. Tongue and H. S. Rzepa, http://www.ch.ic.ac.uk/vchemlab/