(a) Department of Chemistry, Imperial College of Science, Technology and Medicine, London, SW7 2AY and (b) School of Chemistry, University of Leeds, Leeds.
Abstract: A review of the development of computational chemistry tools which make direct use of the Internet is presented, together with some recent advances in new Internet methodologies based around chemical MIME standards, Java Applets, VRML (Virtual Reality Modelling Language) and server-side technologies. The use of these tools in new generations of chemistry electronic journals is illustrated via the CLIC project to enhance the journal "Chemical Communications" and the activities of the "Open Molecule Foundation".
The World-Wide Web allows the 3D coordinates to be integrated into the document describing the research. The molecular content can be rendered on the screen with a chemical MIME aware "plug-in" such as Chime.[4] The syntax of the command that invokes Chime is shown below;
<EMBED SRC="porphend.pdb" bgcolor=#FFFFFF align=abscenter width=250
height=250 spinx=360 startspin=true display3D=sticks name="main"
script="zoom 120; connect true; set bondmode and; set hydrogen
false">
This allows the user to acquire a chemical document from the Internet, and to impose a molecular style on the content such that any 3D molecular coordinates are integrated directly into the screen display. The reader can rotate the 3D representation of the molecule, change its stylistic attributes from the default supplied by the author e.g. wirefame to spacefill, decide which colours to display it in, perhaps measure interatomic distance, even perform computations on the system. Most importantly, they can save all this information either to hard disk, or perform the equivalent of a copy-paste operation from the Web client window to some other program of their own choosing. Thus Chime supports such export/input to a full-featured molecular editor called ISIS/Draw.
The earliest example of how such "hyperactive" molecules were seamlessly integrated into a chemical document is in a keynote paper for the 1995 ECTOC electronic conference.[5] The concept has also been extensively used in the later ECHET96 conference[6] and is now a feature of the electronic version of the Journal "Chemical Communications"[7] for enhancing the keynote articles. In summary, the World-Wide Web, in conjunction with chemical MIME headers, turns a chemical document into a working live tool, where the user makes the decisions on what to do with the data, rather than simply accepting a single point of view imposed by the author or publisher of the information.
Chime operates on two levels, the basic plug-in and an enhanced version called Chime Pro, which will offer the ability to accept a structure query directly from the clipboard, for searching a proprietary database engine such as Chemscape Server. The ability to display data in-line on an HTML page or within an HTML table is promised to give substantial performance benefit for the display of search results from a molecule or reaction database. This refers to the problems with "stateless" communication protocol such as HTTP, in which every component of a document has to be retrieved from a server in a discrete transation. If the user wishes to retrieve say 100 molecules with associated 2D or 3D co-ordinates, then the large number of HTTP transactions required makes the process very inefficient. By retrieving all the molecules in a single transaction, and then parsing the molecules out of this, one achieves much greater efficiency (and one also recovers the "state" or "context" between the molecules). Such databases services, which might in the future be an integral part of reading a research article in a journal represent an impressive technical advance over what was possible even one year ago. This does, however, raise interesting issues of whether it is appropriate to associate the reporting of primary research results with potentially proprietary and commercial information services.
Other aspects of plug-in technology must also be considered. Firstly it is still necessary for the author of the software (in the case of Chime, MDL Information systems) to produce operating system specific versions of the plug-in. For example, MDLI released a Unix version more than six months after the Windows and the Macintosh versions became available, and then only for SGI systems. Secondly, the reader must be pro-active in acquiring this software, and must often download perhaps three different versions to satisfy local installation requirements. Finally, there is also no guarantee that any two chemical plug-ins will necessarily inter-operate, since guidelines for interchange of data between various plug-ins do not appear to currently exist. At least the plug-in implementation by some suppliers, e.g. Netscape V 3.0, now allows the user to choose which plug-in to use for any given supported chemical MIME type, a feature not supported in the initial releases.
We noted above that the Chime Pro plug-in was capable of parsing an HTML document for molecule content, solving the problems of the stateless HTTP protocol. This raises the issue of whether the molecular content should be enclosed in any particular standard form. Ideally, this should follow the strict SGML guidelines from whence HTML originated, but which nowadays seem rarely followed. A secondary issue is whether HTML is the best carrier for molecular content. In recognition of the many inadequacies of HTML as a molecular descriptor, we have continued development of the "chemical markup language" or CML, which was implemented by Murray-Rust[10] and first reported at the 1995 Nimes meeting. CML functions as a medium for the inter-operability of chemical information between areas such as publications, programs, equipment, databases, other structured file formats, for example, CIF, CEX, asn.1 and CXF, and older legacy formats such as PDB. Because it is derived from a formally defined SGML DTD (document type definition), it can be parsed using standard tools. Futhermore because of the highly structured nature of its implementation, it can be associated with the molecular class libraries defined by Java in a natural way. Thus CML encapsulated chemical semantics provide a powerful and extensible way forward for the development of Internet based chemical tools.
By mid 1996, a significant number of molecular Java applets had been produced, including molecular editors, molecule visualisers, sequence alignment editors, front ends to computational programs such as Gaussian 94 and a CML visualiser capable of parsing a CML encoded file [8]. In order provide a mechanism for standards to development in this process, to document the subject in a manner suitable for molecular scientists and to try to ensure future inter-operability is achieved in the area of molecular science, an organisation known as the "Open Molecule Foundation" has been set up to facilities developments in this area, and to provide information and support for developers and users.[9]

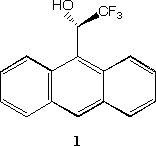
We recently argued[11] that the negative electrostatic potential of 1
is highly chiral, a property that we hypothesised is related to the excellence of
this reagent as a chiral resolving agent via its weak binding and hydrogen bonding properties (Figure 1)
We have previously shown [12] how VRML can be used to present a variety of types of molecular information to the user in an integrated manner. Just as with HTML, VRML objects can contain URL hyperlinks to other Internet based resources. In our case, individual points in the 3D scatter diagrams could be hyperlinked to other VRML scenes containing details of inter-molecular interactions relevant to the chemistry being discussed, and these in turn can be hyperlinked to bibliographic information about each molecule, or whatever. Other possibilities include linking VRML objects to Java applets or scripts which can perform actions on such objects. The most trivial example would be to change the radius of a spacefill display of any particular atom from e.g. the van der Waals value to some other. A more sophisticated example is the use of Java to display digital spectral information derived from an NMR spectrometer,[13] and to link regions of the spectrum to specific atoms or residues in a 3D molecular object displayed using e.g. VRML. The links can be bi-directional, i.e. clicking on a specified atom will highlight the spectral region containing peaks associated with that atom.
Such bi-directional functionality could of course be implemented entirely using Java applets, and as the 3D rendering tools available in Java improve, this route may well become the favoured one. Indeed with the Java 3D API class now under development, there is no reason why a VRML viewer cannot be completely written in Java. Such a Java-based VRML viewer would consolidate the two technologies and preclude the need for any VRML plug-in. Certainly, the current trend towards the production of very memory intensive Web browsers to which the user might have added a number of plug-ins and potentially duplicate functionality seems unsustainable.
The future of VRML itself depends on the quality of the content available. Tools, such as EyeChem 2.0, [12] are required to facilitate the creation of molecular VRML files from scratch or from data in other formats. A drawback of VRML is that this file format is ill suited for molecular data. One approach to this problem would be to prototype a more appropriate 3D file format, such as the Molecular Inventor [14] format into VRML. Another approach to this problem would be to implement an SGML document type representation for VRML [15] which would also address indexing and archiving issues of VRML files in a structured document server.
As described thus far, HyperWave has no explicit molecular "intelligence". However, in the future it is expected that this will be added by installing other SGML DTDs to the server functionality, such as for example the CML DTD, or enhanced versions of the ISO 12083 DTD which contain chemically explicit entities. Experiments underway will establish whether this approach will offer the type of functionality and in particular the performance suitable for chemical databases.
A second theme addresses the lack of structure that is already apparent in the so-called CGI (Common Gateway Interface) mechanisms for processing subject specific content. A "cgi" program or script is frequently used to perform specific tasks based on user-provided information derived from a forms-based input, and to return the output to the user via the browser window. These custom written "cgi" processes suffer considerably from a lack of explicit guidelines, and over a period of time they can become essentially unmaintainable. Such scripts are often also not inter-operable; frequently it is easier to introduce minor variations via a new script than to modify the existing script. Whilst this is currently a largely unsolved problem for chemistry, new approaches to the integration of the Web server with its "cgi" functionallity are now appearing. Both the Jigsaw server from the World-Wide Web Consortium and the Jeeves server from Sunsoft offer a Java-based solution. In this scenario, the "cgi" functionality is closely integrated in an object-oriented manner to the server itself, thus offering a stable and scaleable way of building discrete subject-based services into the document collection. A solution which offers chemical functionality via both the server and the document structure seems promising development path to follow.
Acknowledgements: The authors thank Peter Murray-Rust, Jurgen Brickmann and Adam Precious for their inspiration and collaboration. Funding from the JISC Electronic libraries (e-Lib) and JTAP programs, from British Telecom for a University Development award and from GlaxoWellcome is gratefully acknowledged.
[2] B. J. Whitaker and H. S. Rzepa, "Chemical Publishing on the Internet", Conference on Chemical Information, Nimes, France, October, 1995 (http://www.chem.leeds.ac.uk/papers/html/Nimes/nimes.html). See also http://www.ch.ic.ac.uk/chemime/iupac.html for the latest summary of this area. [3] For the official chemical MIME home page, see http://www.ch.ic.ac.uk/chemime/
[4] See the URL http://www.mdli.com/ Chime is itself a Netscape plug-in enhancement of the RasMol molecular viewer written by Roger Sayle. For the history of RasMol, see http://www.glaxowellcome.co.uk/netscape/software/history.html
[5] A. Padwa, E. A. Curtis, V. P. Sandanayaka, and M. Weingarten, ECTOC, 1995, See http://www.ch.ic.ac.uk/ectoc/papers/01/
[6] See http://www.ch.ic.ac.uk/ectoc/echet96/
[7] The CLIC Project. See http://chemcomm.clic.ac.uk/
[8] For one such collection, see http://www.ch.ic.ac.uk/java/
[9] For information about Java and tools for Bio and Chemo-informatics, see the Open Molecule Foundation; http://www.ch.ic.ac.uk/omf/
[10] P. Murray-Rust; http://www.dl.ac.uk/CBMT/cml/cml06f/newintro/role.html
[11] For further details, see D. O'Hagan and H. S. Rzepa, Chem. Commun, 1996, in press. An electronic version of this article will also be available via the journal home page; http://chemcomm.clic.ac.uk/
[12] For chemically oriented examples of how VRML has been applied, see O. Casher and H. S. Rzepa, J. Mol. Graphics,1995, 13, 268; H. Vollhard, C. Henn, G. Moeckel, M. Teschner, J. Brickmann J. Mol. Graphics, 1995, 13, 368; J. Brickmann, H. Vollhardt, Trends In Biotechnology, 1996, 14, 167-172.
[13] H. S. Rzepa, P. Murray-Rust and R.Kinder; See http://www.ch.ic.ac.uk/java/HyperSpec/
[14] O. Casher and H. S. Rzepa, Proceedings of the 14th UK Eurographics Conference, March, 1996. See http://www.ch.ic.ac.uk/rzepa/eg/
[15] W. E. Kimber; See http://38.145.245.206:80/drmacro/vrml/