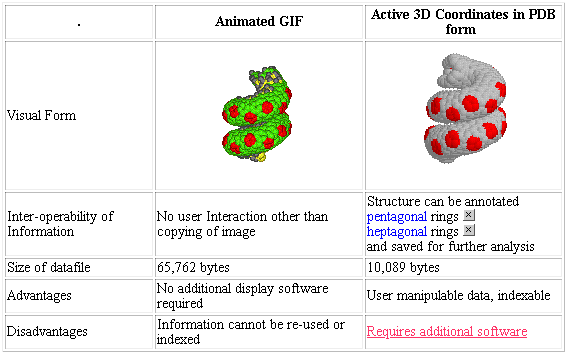
Figure 1. Comparison of Molecular Structures expressed in Bit-mapped Image and in PDB Form.
(If you are viewing the on-line version of this article, click on the image to get an active form of this table)
Chemists were amongst the earliest users of computers for generating, acquiring or searching for data, and hence deriving information in order to create knowledge about the subject.1 The established access model dating from the early 1980s is the use of proprietary on-line networks such as STN (Scientific and Technical Network). An alternative global information paradigm based on the Internet and known as the World-Wide Web was introduced in the early 1990s, and has subsequently gained widespread general use. This review will introduce some of the innovative and useful features that characterise the Web and in particular its chemical content, how it can be used to create new connections between related subject disciplines, and how it has the potential to influence the manner of future scientific collaboration.
The review is not intended as a technical handbook. Nor in an area where the speed of change and growth is unusually high, can we attempt in print to be either comprehensive or entirely current. In keeping with the subject of this review, an electronic version of the article is available2 for those readers who wish a starting point for their exploration of the chemical themes explored here, or who wish to investigate those aspects of this article which cannot be demonstrated in print.
The seamless integration of on-line chemical and biological data archives with other areas of scientific interaction such as journal publishing, conferences, electronic mail and discussions, books and other reference sources, laboratory systems, commercial catalogues, computer based research activities, and human resources could be described as something of a holy grail in science. The potential to achieve this ideal is thought to have come closer with the introduction in 1989 by Berners-Lee and colleagues3 of the World-Wide Web system, and its subsequent rapid adoption from around 1994 onwards by sections of the scientific and chemical communities.
To help highlight why the Web differs radically from earlier infra-structures, the characteristics of the more traditional on-line information model are first summarised. Access is usually controlled by individual user accounts via a small number of remote server access points for users, and requires use of custom software which tends not to be inter-operable between different vendors. With a few exceptions, mechanisms for individuals or local organisations to innovate with new components in a scaleable and integrable manner are not made available. In essence, the models have tended to be monolithic and centralised rather than collaborative and distributed. Local "points-of-access" to such services have traditionally been located in libraries, and other specially designated areas. It is only in the last five years or so that the infra-structure has become available in most organisations for extending this access to offices, laboratories, meeting rooms, lecture theatres, the home, and indeed travel between these locations if desired.
The use of the word "resource" implies that not only documents, but
other real or potential sources of data and information can be
uniquely located on a global scale. In a chemical context, this could
include database search engines and other computational algorithms and
resources, instruments, people or scholarly articles. Even more
importantly, quite finely grained data within a document might be
addressed, as for example individual atoms in a set of molecular
coordinates, or peaks in a spectral representation. A typical example
of a URL might be the one associated with the article you are
currently reading;
http://chemcomm.clic.ac.uk/csr/rmw.html#intro
The first part of the string indicates the use of Hyper-Text Transport
Protocol (HTTP), a mechanism used to exchange information between the
computer where the resource resides (the server) and the computer
which the user is using to acquire the information (the client). HTTP
has one relevant characteristic; it is said to be
"stateless". Thus a HTTP transaction between a server and a
client is entirely self contained, typically lasting perhaps a few
seconds in duration, and there is no state or context maintained
between any two HTTP transactions. This feature, which was originally
chosen because of its excellent network characteristics, differs
fundamentally from the type of context-rich environment associated
with e.g., CAS on-line or other dedicated database systems.
Instead on the Web, context is currently achieved predominantly
via a device known as the hyperlink (vide
infra), although there is some discussion that the stateless HTTP
protocol might be replaced in future by a richer "stateful"
system.
The component chemcomm.clic.ac.uk specifies the precise network device where the information resource will be found, being mapped onto one IP address of the type mentioned above. The final component /csr/rmw.html#intro specifies a hierarchical directory structure, a file name with an associated extension and a named "anchor" (#intro) within the file. The anchor in this case refers to the introduction in this article, but it could just as easily refer to e.g. an individual atom in a molecular descriptor. The file name extension .html indicates a document written in a descriptive language called HTML (Hyper-Text Markup Language).
<command>Text or other content acted upon </command>
where "<" and ">" are special characters describing an HTML command, and where <command> and </command> define the start and end of the content acted upon by the command. The choice of a markup language for the Web is significant. Because a distinction is made between the content of a document or resource and the manner in which it is displayed, this latter task is performed less by the author of the resource, and more by the client program used to display it. This program is often referred to as a "browser", and the user of the program can specify, for example, the style, size, and colour of any displayed text depending on the device they are using to view the data, or in a chemical context how a molecule might be displayed.
Because of the focus on information content, documents written in HTML are relatively easy to index, at least in a non-chemical sense. Within two years of the increasing use of HTML to encode content on the World-Wide Web, software had been developed that allowed the global Internet to be indexed and hence searched. At the end of 1996, some 80 million documents had been so treated. However, because HTML is predominantly a generic descriptive language based around text, it does not define any specifically chemical descriptors. For this reason, other means of expressing this subject have been hitherto been adopted. The earliest and still the most common solution was to use bit-mapped graphical images in so-called GIF (Graphical Interchange Format) or JPEG (Joint Photographic Experts Group) formats. Other than to a human being, these are largely "content free" and almost un-indexable (e.g. Figure 1). Inevitably, as the World-Wide Web has become more commercially oriented with an increasing focus on style and "look and feel", the use of bit-mapped images has proliferated, and this single aspect is causing concern as the world's networks become overloaded with their transmission.
Unlike a service such as STN, there is no single unambiguous entry point for users wishing make productive use of the World-Wide Web. Instead, there are a variety of mechanisms available, some of which are genuinely global in nature, others of which depend entirely on the local characteristics of the users environment, and many of which carry no cost to the user. Which one is used depends very much on the nature of the problem being addressed.
Because of the focus on hyperlinked contextual content rather than style in the original specification of the HTML, indexing the collective content on a global scale has proved a productive, and indeed commercially lucrative operation. Because HTML is not particularly appropriate for encoding specifically chemical information, one would use these global indices when searching for more specific concepts that are easily defined using simple keyword terms. Collected in Table 1 are some of the more popular indexed collections available in 1996, along with the results of a search based on using "chemistry" or "chemical" as keyword terms. Particularly noteworthy is the HotBot resource, which allows searches to be performed on files other than HTML which may be better suited for encoding chemical content (e.g. 3D coordinate files in the PDB format).
| Search Engine | URL | Hits for "Chemistry" or "Chemical" (September 1996) |
|---|---|---|
| Lycos | http://www.lycos.com | 17,094 |
| Alta Vista | http://www.altavista.digital.com/ | "About 600,000" |
| HotBot | http://www.hotbot.com/ | 1,053,081 |
| InfoSeek | http://ultra.infoseek.com/ | 302,667 |
| Yahoo Chemistry | http://www.yahoo.com/Science/Chemistry/ | 26,464 |
| Chemical Collection | URL |
|---|---|
| Virtual Chemistry Library Collection | http://www.chem.ucla.edu/chempointers.html |
| ChemDex at Sheffield | http://www.shef.ac.uk/uni/academic/A-C/chem/chemistry-www-sites.html |
| The RSC Collection | http://chemistry.rsc.org/rsc/ |
| The ACS ChemCenter Collection | http://www.ChemCenter.org/ |
| ChemWeb Collection | http://chemweb.com |
| Entry | Information Source | URL |
|---|---|---|
| 1 | The Brookhaven Protein Databank | http://pdb.pdb.bnl.gov/cgi-bin/browse |
| 2 | The Cn3D-Entrez System | http://www.ncbi.nlm.nih.gov/Structure/cn3d.html |
| 3 | MDLI | http://www.mdli.com |
| 4 | Daylight Information Systems | http://www.daylight.com |
| 5 | Tripos Sketch & Fetch | http://www.tripos.com/SandF.html |
| 6 | The WWW Chemical Structures Database5 | http://schiele.organik.uni-erlangen.de/services/webmol.html |
| 7 | The ChemFinder System6 | http://chemfinder.camsci.com |
The WWW Chemical Structures5 and the ChemFinder6 databases are noteworthy because they are based in part on molecular information retrieved from the Internet itself. The ChemFinder database from CambridgeSoft is primarily a text-based index of a manually selected number of database sites containing CAS registry number, compound names, connectivity and physical data. The principal emphasis is on checking the compound names extracted from the indexed HTML pages and automatically correcting any errors. In contrast, the WWW Chemical Structures database was constructed from the results of an index robot designed for the task of finding 2D connection tables and 3D molecular coordinates on the Internet. In excess of 2200 such entries were retrieved from the Internet as it existed in 1995. The construction of these databases involved a significant degree of manual error correction because the chemical content was not always created in a consistent and standard way by the original authors. That it was possible at all is because some standards do exist for this purpose.
If one compares the results of global keyword (Table 3) and molecular structure searches5, a ratio of one discrete structure was found for every 100+ Internet based documents which contain the word "chemistry". This is almost certainly because most molecular structures are currently represented on the Internet by bit-mapped GIF or JPEG images, from which the semantic molecular content can be extracted only with great difficulty, and which therefore cannot be automatically indexed in any chemical sense. The following example (Figure 1) compares the characteristics of a representation of a helical carbon nanotube as an animated GIF file and as a PDB (Protein DataBank) file which has been rendered into visual form on the computer screen.

In order to explain the distinction between the two molecular representations, we have to explain the underlying standards and software technologies that have been introduced in the last three years7.
A large variety of file formats are currently utilized for storing electronic information. This is acceptable when running proprietary tools on dedicated hardware but begins to be problematic for information designed to be shared over a network comprising of many platforms and operating systems. In 1993, Borenstein and Freed8 proposed a mechanism called MIME (Multipurpose Internet Mail Extensions) which would allow a variety of standard file formats to be exchanged over the Internet using electronic mail. It works by the simple device of adding a short header at the top of each datafile attached to a mail message which provides the information needed for the mail handler to process the contents. The header is hierarchical, with a primary designation designed to provide some measure of sensible "default" handling of the content, and a sub-type which is more specific. So for example, an HTML file is indicated by text/html as a so-called media type.
The mechanism was rapidly adopted for use with World-Wide Web clients. When a user makes a selection through a hyperlink within a HTML document, the client browser "posts" the request to the designated web server. Assuming the server accepts the request, it locates the appropriate file(s) and sends them to the client, with the relevant MIME header attached. When the browser receives the data, it reads the MIME type to determine what to do with it. For MIME types such as text/html or image/gif the browsers have been built in such a way that they can simply display the information in the browser window. For other MIME types, a local preference file is inspected to determine what (if any) local program (known as a helper application) can display the information, this program is then launched with the data file and the result displayed in a newly opened application window. The important aspect of this mechanism is that it achieves the delivery of semantic content to the user, who can specify the style in which it will be displayed via their choice of an appropriate application program.
In early 1994, we proposed7 a coherent set of standard chemical file types and made a proposal for the introduction of what we called the Chemical MIME standard. A number of different chemical media types were originally proposed,9 falling into several categories; (1) molecular coordinate and connectivity and sequence formats, (2) molecular modelling formats, (3) spectroscopic formats (4) generic "self-defining" modern formats. A collection of representative types is shown in Table 4; a comprehensive and current list is available on-line.9 The open architecture allows for new MIME types to be proposed and adopted without having to create new servers or clients. However, we envisage a small core of fundamental chemical MIME types passing through a formal ratification process, whilst new types of perhaps a more proprietary nature of interest to chemical software houses remain as "x" types, although nevertheless registered via a central mechanism for coordinating and documenting these types.
| chemical/x-pdba | Brookhaven Protein data base format |
| chemical/x-daylight-smiles | Daylight chemical connectivity format |
| chemical/x-mdl-molfile | Molecular and Reaction formats defined by MDLI |
| chemical/x-mdl-tgf | Transportable Graphic format defined by MDLI |
| chemical/x-c3d | 3D Format defined by CambridgeSoft |
| chemical/x-embl-dl-nucleotide | SwissProt data format |
| chemical/x-ncbi-asn1 | Protein Sequence Format |
| chemical/x-gcg8-sequence | Protein Sequence Format |
| chemical/x-kinemage | Protein Cartoon Format |
| chemical/x-csml | Chemical Structure Markup Language |
| chemical/x-mopac-input | Mopac Input Format |
| chemical/x-gaussian-input | Gaussian Input format |
| chemical/x-jcamp-dx | Standard for defining MS, IR and NMR spectra |
| chemical/x-cif | Crystallographic Interchange format |
| chemical/x-cxf | Chemical eXchange Format |
| chemical/x-cml | Chemical Markup Language |
The advantage of using chemical MIME as a simple descriptor of molecular information is twofold. Firstly, at least for molecules containing fewer than around 500-1000 atoms, the data file describing the atomic co-ordinates is smaller than a bit-map image at reasonable resolution and colour depth of the molecular structure. Thus when the data are downloaded from the server to the client, less precious network bandwidth is used. For example the file size needed to describe a helical carbon nanotube as an animated GIF is over 6.5 times larger than the co-ordinate file (Figure 1). Secondly the quality of the information transferred is of a higher value. Once the atomic co-ordinates are known, the reader can manipulate the image in the browser to rotate the molecule, view it from a different perspective or change the manner in which the structure is presented, e.g. as a spacefilled model of a small molecule or a ribbon cartoon of the backbone structure of a protein.
Because the molecule is described in terms of atoms and their
coordinates, even individual functional groups can be addressed. This
was first achieved using a mechanism called Chemical Structure Markup
Language (CSML)7, which comprised small scripts held on
the Web server and identified with the chemical MIME type
chemical/x-csml
This enables individual regions of molecular structures to be
addressed from hyperlinks embedded in text files or graphical images.
If you are reading the "active" version of this article on
the Web with the Chime molecular viewing plug-in installed, you can
test this for yourself by clicking on the buttons in Figure 1 labelled "pentagonal" and
"hexagonal" rings respectively, This highlights the
functionality which induces the carbon nanotube to adopt a helical
conformation. CSML is now regarded as the precursor to the later
development of a properly structured Chemical Markup Language (CML,
see below) based on SGML guidelines13.
The availability of software supporting chemical MIME types has enabled the rapid development of many innovative applications, of which just a few examples14 include medical biochemistry tutorials,14a correlating cross-peaks in 2D spectral data with atom pairs,14b the development of interactive workshops in which students can explore the structural properties of inhibitors specific to HIV protease,14c an annotation of the activity of taxol14d and a guided tour through the photosystem reaction centre highlighting the course of electron transport.14e The adoption of MIME standards can be viewed as the first key event in the development of chemical publication on the Internet by introducing a new paradigm for the publishing of a rich variety of chemical data.15
The methods covered so far relate to the visualisation and interpretation of a relatively small range of molecular data. However, the variety of disciplines and techniques that chemistry covers is enormous, so it is not surprising that information exchange between different types of molecular data is difficult. The traditional approach has been either to try to standardise on a single format (e.g. chemical/x-pdb for proteins, and increasingly for small molecules also) or to write conversion programs such as Babel16. Unfortunately the latter process always implies information loss: for example the MDLI Molfile format does not hold bibliographic information and in turn PDB files do not hold full connection tables.
A more serious problem is that electronic information decays. The formats used today may be (literally) indecipherable in 5 years time; many do not even have formally published standards but rely on word-of-mouth and guesswork. Even when manuals are available, it is often difficult to know whether two developers apply the same semantics to a given term. It is generally accepted that the best way to tackle these problems is through the use of markup languages and public discussion. Markup languages add so-called meta-information to a document to allow it to be processed in a contextually rich manner.
<!DOCTYPE article PUBLIC "-//CSR/DTD article/EN">
<ARTICLE>
<TITLE>The World-Wide Web as a Chemical Information Tool</TITLE>
<ADDRESS>Department of Chemistry, ... </ADDRESS>
<AUTHOR>Peter Murray-Rust, Henry S. Rzepa
and Benjamin J. Whitaker</AUTHOR>
<ABSTRACT> Chemists were amongst the earliest
users of computers ...
</ABSTRACT>
<SECTION>
<SECTIONHEADING>The
Characteristic Features of The World-Wide Web </SECTIONHEADING>
The information model ...
</SECTION> ...
</ARTICLE>
The first line contains a document type declaration identifying the DTD required for the document - in this case a (fictitious) declaration for a document in the style of an English language (EN) article for Chemical Society Reviews. This is followed by a start tag <ARTICLE>, identical to the tag used for the DTD declaration, which is called the base document element and which activates the DTD. The elements following the <ARTICLE> start tag are fairly self-explanatory. Finally the end of the document is identified with a closing end tag </ARTICLE>. Notice that only the logical structure of the document is important, for example the <ADDRESS> ... </ADDRESS> container appears out of sequence from that in the printed version of this article. Textual elements may be nested so, for example, we could define a container <CITATION>...</CITATION> with sub elements <AUTHOR>...</AUTHOR>, <YEAR>...</YEAR>, <PAGES>...</PAGES>, and so on. SGML tags may be also qualified by attributes as in <CITATION type=Book> which might make certain sub elements mandatory, e.g.<PUBLISHER>. Each SGML document, known as an SGML document entity, consists of three components, an SGML declaration defining the syntax and character set used within the document, the DTD defining the logical structure of the document, and the text itself.
These are quite general, so that markup might appear as:
<X.VAR TITLE="Heat of Formation", REL="glossary"
HREF="/chem/theor?deltaHform" UNITS=kilocalorie/mole">12.34</X.VAR>
or
<C.MOL TITLE="1,3,5-cyclohexatriene", REL="glossary"
HREF="/cml/organic?benzene" </C.MOL>
CML documents can have a very flexible structure which allows a large and finely grained body of chemical information to be encoded, such as;
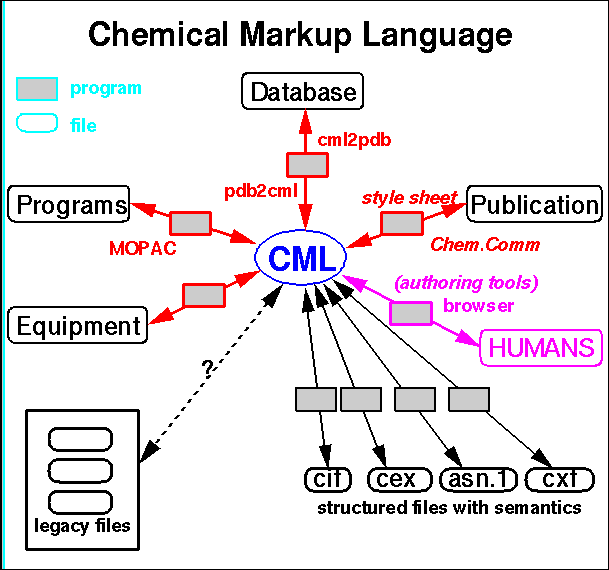
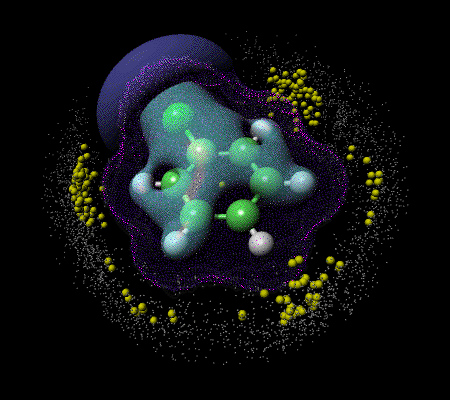
A number of interesting chemical applications of VRML have been published in the last two years 19. The earliest use was by Casher and co-workers to demonstrate how a so-called molecular collaboratory could be constructed19a. This involved the use of high speed national computer networks combined with videoconferencing techniques to allow two or more groups of collaborators to simultaneously explore complex molecules. Associated properties such as computed molecular orbitals or electrostatic potentials are also well handled by this technique.19b Vollhardt and Brickmann19c have used VRML extensively to describe other complex molecular surfaces and properties including active site highlighting, and Robinson and Hardy19d have applied it inter alia to cell membrane model construction. The use of VRML is particularly appropriate where the context between a set of diverse three dimensional properties needs to be analysed. For example, clustering analysis of intermolecular interactions derived from crystallographic information can be integrated with molecular diagrams, computed electronic properties and bibliographic information about individual molecules19e (Figure 3).
The Java language allows the creation of a stand-alone application, but more interestingly of a network-portable version called an applet. An applet is downloaded to the client upon request and executed locally within a Web browser. Unlike browser plug-ins, Java code is a meta-language and hence platform independent, requiring only a Java-compliant local environment. This can be a Web client such as Netscape or it can be a feature of the operating system itself. This eliminates the often complex step of a user having to install a new program on their local computer system. If a document written in a language such as CML is being viewed, only those Java applets that relate to the document content need be acquired. If the reader wishes to view an infra-red spectrum for example, only the Java objects that can read and display such data need be acquired.
Java applets are relatively new technology. Their immediate appeal has resulted in an explosion of interest and development, and in late 1996 there were already a number of excellent applets freely available on the Web,21 including sketching applications which allow structures to be defined in database queries (Table 3). An organisation called the Open Molecule Foundation22 has been established to serve as one focal point for identifying, resourcing and coordinating development in this area.
In this final section, we concentrate on two areas of scientific collaboration and dissemination which have been significantly influenced by the technologies and infra-structures described above. There are of course many other areas which space does not permit us to discuss, including the impact of the Web on teaching practice, the development of commercial models in this environment and aspects such as copyright, privacy and data protection, peer-review and authentication.
In November 1994, the first entirely Web-based chemical conferences, one on chemometrics and the other on computational chemistry were organised (Table 5). Their description as a "conference" seems to have persisted, although it seems best to regard such events as representing something of a new genre for scientific exchange. The novel aspect of these conferences was their integrated use of various Internet based technologies such as thematic electronic mail, conference articles presented as HTML documents, and the use of a "MOO" (Multi-user Object Oriented) text based interactive discussion environment. ECCC-123 was the first to adopt the use of chemical MIME to present 3D molecular models to the reader, and indeed was the first such conference where the proceedings were subsequently published on CD-ROM and recognised by Chemical Abstracts. This model was also followed in 1995 by the ECTOC-1 conference on organic chemistry,24 which introduced keynote articles, full text based keyword searching and an integrated molecular hyperglossary which could be searched by chemical sub-structure. Participants could also look at the access statistics to the conference to find out which the most popular articles were, and submit to a conference photographic collage. The second conference in this series (ECHET96) further developed the genre by allowing authors to edit their own contributions interactively via a Web interface, and by the use of molecular similarity clustering,25 offering suggestions to participants on papers that might be related to the one they were reading. Due possibly to their relative novelty, these early conferences accumulated impressive access statistics, typically involving in excess of ten thousand visitors in a 1-2 month interval.
Events such as the Electronic Glycoscience conference (ECG-1) evolved a slightly different metaphor, in which the various themes of the conference were mapped onto an imaginary location analogous to a real conference venue. Participants had to register (at that time for no fee) before they could browse, and interactive MOOs were used extensively for real time discussions of individual contributions and other themes. Unlike previous e-conferences, the ECG-1 proceedings were subsequently published in a conventional printed journal, and this trend has been continued with subsequent events by the same organisers (ECG-2, e-MGMS).
| Conference | URL | Published Proceedings |
|---|---|---|
| InCINC'94: Chemometrics | http://www.emsl.pnl.gov:2080/docs/ incinc/homepage.html | Various Journals |
| ECCC-1: Computational Chemistry | No longer available | S. M. Bachrach, W. Hase, D. B. Boyd, H. S. Rzepa, S. K. Gray, (Eds), Proceedings of the First Electronic Computational Chemistry Conference, (ARInternet Corp., Landover, Md.), 1995. |
| ECTOC-1: Organic Chemistry | http://www.ch.ic.ac.uk/ectoc/ | H. S. Rzepa, C. Leach and J. M. Goodman, "Electronic Conference on Trends in Organic Chemistry", Royal Society of Chemistry, ISBN 0 85404 899 5, 1996. |
| ECG-1: Glycoscience | http://bellatrix.pcl.ox.ac.uk/ egc/Welcome.html | B. H. Wilson, B. J. Hardy, Tr. Glycoscience Glycotechnology, 1996, 8, 301. Proceedings in Glycoconjugate J.. |
| TMMec: Molecular Modelling | http://uqbar.ncifcrf.gov/ agora/welcome.html | Various Journals. |
| ECCC-2: Computational Chemistry | http://hackberry.chem.niu.edu/ECCC2/ | Theochem |
| ECHET96: Heterocyclic Chemistry | http://www.ch.ic.ac.uk/ectoc/echet96 | H. S. Rzepa, C. J. Moody, R. Jones, A. Padwa, J. Snyder and C. Leach, (Eds), "Electronic Conference Heterocyclic Chemistry", Royal Society of Chemistry, 1997, to be published. |
| e-MGMS: Molecular Modelling and Graphics | http://bellatrix.pcl.ox.ac.uk/ mgms/Welcome.html | J. Mol. Graphics. |
The preceeding examples were of conferences conducted exclusively on the Internet. However such electronic enhancement has had a valuable role as an adjunct to physical events, and since 1995, many international scientific conferences have had offered a Web component.26 This can range from making available before the start an electronic book of abstracts, an electronic registration desk at which participants could supply their personal details, and obtain travel and tourist information.
In late 1996, it seems likely that several models for electronic conferences will continue to evolve. All will continue to offer new forms of "added value" functionality, some will be purely evanescent, with authors having the option of subsequently submitting their contribution to more conventional journals, others seem likely to operate on a commercial basis with registration fees being levied. Perhaps the ultimate future of such events is as part of a more complete information club in which the distinction between journals, conferences and informal discussion becomes progressively more blurred.
There are many issues to consider when applying an electronic medium to a scientific journal, from the perspective of the publisher, the author and the reader alike. The opportunities for all are obvious. From the publishers' point of view, to name but a few, there are opportunities to speed up the process of publication, to create "value added" products and to monitor document access and usage by readers. From the author's and the reader's perspective, the attraction lies in the novel ways in which data can be presented and interacted with. More fundamentally, the Web enables access to a wide variety of information on-demand, conveniently and efficiently. The drawbacks of the electronic medium can in the main be put down to lack of precedent. The technology is relatively new and is developing so rapidly that there has simply not been time to legislate for problems such as copyright, privacy and data protection. However, the obvious promise that this medium offers has led to a number of projects experimenting with the possibilities of electronic publication. These can be broadly divided into two types, those which derive from printed journals and which strive for electronic delivery with a degree of faithfullness to the printed form, and those where the prime focus is to enhance the journal concept without the constraint of requiring a precise printed equivalent.23 Whilst experiments in the former category have been conducted for more than twenty years, the latter is a relatively new phenomenon. Here we will focus on one such project of this type, the CLIC consortium initiative15 to establish an electronic version of Chemical Communications, published by the Royal Society of Chemistry.
Although mainly used by structural and organic chemists Chem. Comm. is a general chemistry journal which aims to disseminate rapidly internationally important research results. Each issue also contains a longer feature article highlighting the work of a leading group. Whilst developing a strategy for the implementation of an electronic publication, several issues have arisen. These include;
The solution adopted in the first two areas by the RSC, along with many other publishing houses, has been to make use of SGML derived technologies. This is a complex technical area, and a detailed discussion of this is beyond the scope of the present review.28 Instead we will focus on the some of the "added value" aspects that the use of the World-Wide Web enables. Examples of added value could be to embed interactive 3-D molecular viewers, using VRML or a mechanisms based on MIME types to automatically launch external applications when a piece of embedded data is accessed by the user. NMR spectral data, reaction schemes and equations can also be treated similarly. We have called this concept "hyperactive molecules" and demonstrated it in a number of applications7.
Indeed, one can envisage any object in an article becoming an interactive feature, e.g. equations could be dynamically linked to graphing or symbolic algebra tools. Other possible features of an electronic journal include searching and retrieving data on-the-fly, indexing, hyperglossaries, keyword searching and forward referencing. All but the latter are self evident - forward referencing is the concept that a document in the archive can be dynamically updated whenever a subsequent document refers back to it so generating a hypertextual link 'forward in time'.
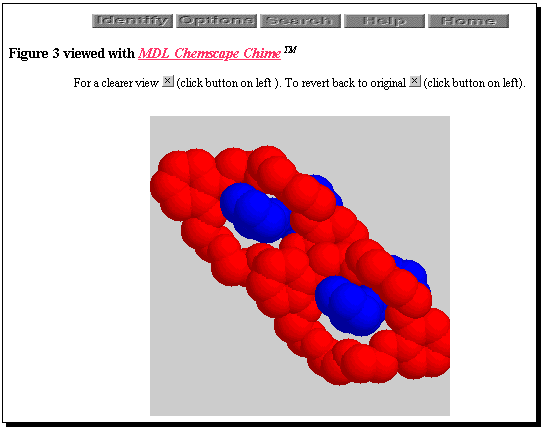
Examples of what can be achieved29 are the enhanced versions of feature articles taken from Chem. Comm. The paper by Stoddart and co-workers30 describes recent advances in the area of the molecular design of interlocking molecules. In the electronic version figures in the paper version depicting molecular structures become animated 3-dimensional models using the Chemscape Chime plug-in. Figure 4 depicts this feature of the article as it appears on the readers screen. Other features in the demonstration article include hypertextual markup of spectral information and extensive cross-referencing.
| Term | Brief Description |
|---|---|
| Bookmark | A user pointer to a previously visited World-Wide Web document |
| Browser | A Client program for viewing World-Wide Web documents |
| CERN | European Laboratory for Particle Physics |
| CGI | Common Gateway Interface for scripts and other external programs |
| CML | Chemical Markup Language |
| DTD | Document Type Description |
| FORM | A graphical interface for interacting with WWW services |
| GIF | Graphical Interchange format (Compuserve) |
| Helper | An independent program invoked from a WWW browser |
| HTML | Hypertext-markup-language |
| HTTP | Hypertext-transport-protocol |
| Hyperglossary | A glossary of terms created and accessed via the WWW |
| Hyperlink | A pointer within a document to another network object |
| Inline | An image inserted within text and displayed by a WWW browser |
| IP | Internet Protocol, currently Version 4, but migrating to Version 6 |
| Java | An object oriented language with characteristics suited for the Internet |
| Javascript | A scripting language with some characteristics related to Java |
| JPEG | Joint Photographic Experts Group image format |
| MIME | Multipurpose Internet Mail Extension, used as two-level descriptor for data files |
| PHP | Personal home page on the World-Wide Web |
| Plug-in | A small program which enhances the capability of some Browsers |
| Server | A Program providing access to a database of WWW documents |
| SGML | Standard Generalised Markup Language |
| URL | Uniform Resource Locator, used within HTML to located hyperlinks |
| VRML | Virtual Reality Modelling Language, a 3D scene descriptive language |
| Web | World-Wide Web |
| Webmaster | The administrator of a World-Wide Web server |
| WWW | World-Wide Web |
Henry S. Rzepa was born in London in 1950, and received both his BSc and PhD degrees from Imperial College, in 1971 and 1974. Following a post-doctoral period with Michael Dewar at the University of Texas, Austin, he returned to Imperial College, where he is now a Reader in Organic Chemistry. His research interests include the theoretical study of stereoelectronic effects, structural studies of unusual forms of hydrogen bonding such as p-facial interactions and those responsible for chiral recognition, and the development of the Internet as a chemical tool. He is the recipient of the 1995 Löschmidt Prize for physical organic chemistry.
Dr. Benjamin Whitaker is a graduate in Chemical Physics from Sussex University, where he also obtained his DPhil, in 1981, for work in laser spectroscopy. His main research interests are in quantum resolved studies of elementary reactions using molecular beams and in the development of laser spectroscopy, particularly degenerate four wave mixing and cavity ring down spectroscopy. He is currently a Senior Lecturer at the University of Leeds.